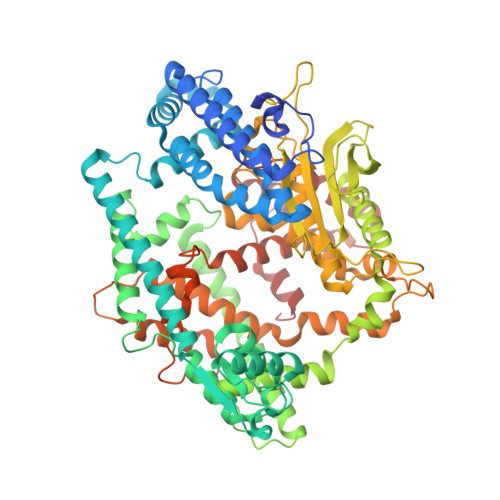
Structure of the Arabidopsis thaliana TOP2 oligopeptidase.
Wang, R., Rajagopalan, K., Sadre-Bazzaz, K., Moreau, M., Klessig, D.F., Tong, L.(2014) Acta Crystallogr Sect F Struct Biol Cryst Commun 70: 555-559
- PubMed: 24817709 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X14006128
- Primary Citation Related Structures:
4PUT - PubMed Abstract:
Thimet oligopeptidase (TOP) is a zinc-dependent metallopeptidase. Recent studies suggest that Arabidopsis thaliana TOP1 and TOP2 are targets for salicylic acid (SA) binding and participate in SA-mediated plant innate immunity. The crystal structure of A. thaliana TOP2 has been determined at 3.0 Å resolution. Comparisons to the structure of human TOP revealed good overall structural conservation, especially in the active-site region, despite their weak sequence conservation. The protein sample was incubated with the photo-activated SA analog 4-azido-SA and exposed to UV irradiation before crystallization. However, there was no conclusive evidence for the binding of SA based on the X-ray diffraction data. Further studies are needed to elucidate the molecular mechanism of how SA regulates the activity of A. thaliana TOP1 and TOP2.
- Department of Biological Sciences, Columbia University, New York, NY 10027, USA.
Organizational Affiliation: