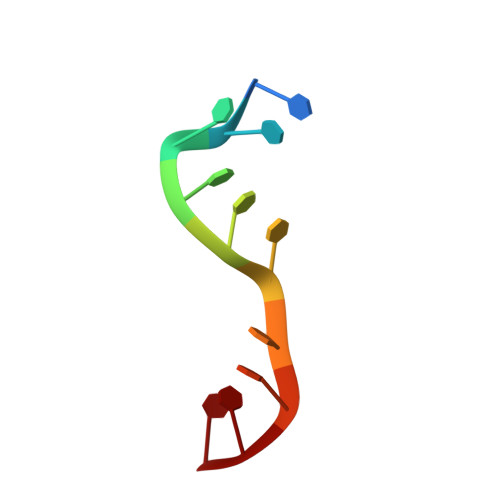
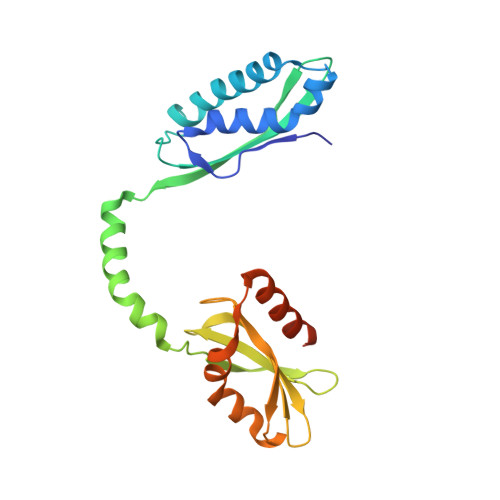
Entrapment of DNA in an intersubunit tunnel system of a single-stranded DNA-binding protein.
Ghalei, H., Moeller, H.v., Eppers, D., Sohmen, D., Wilson, D.N., Loll, B., Wahl, M.C.(2014) Nucleic Acids Res 42: 6698-6708
- PubMed: 24744237 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gku259
- Primary Citation Related Structures:
4PSL, 4PSM, 4PSN, 4PSO - PubMed Abstract:
Instead of a classical single-stranded deoxyribonuleic acid (DNA)-binding protein (SSB), some hyperthermophilic crenarchaea harbor a non-canonical SSB termed ThermoDBP. Two related but poorly characterized groups of proteins, which share the ThermoDBP N-terminal DNA-binding domain, have a broader phylogenetic distribution and co-exist with ThermoDBPs and/or other SSBs. We have investigated the nucleic acid binding properties and crystal structures of representatives of these groups of ThermoDBP-related proteins (ThermoDBP-RPs) 1 and 2. ThermoDBP-RP 1 and 2 oligomerize by different mechanisms and only ThermoDBP-RP2 exhibits strong single-stranded DNA affinity in vitro. A crystal structure of ThermoDBP-RP2 in complex with DNA reveals how the NTD common to ThermoDBPs and ThermoDBP-RPs can contact the nucleic acid in a manner that allows a symmetric homotetrameric protein complex to bind single-stranded DNA molecules asymmetrically. While single-stranded DNA wraps around the surface or binds along channels of previously investigated SSBs, it traverses an internal, intersubunit tunnel system of a ThermoDBP-RP2 tetramer. Our results indicate that some archaea have acquired special SSBs for genome maintenance in particularly challenging environments.
- Freie Universität Berlin, Fachbereich Biologie, Chemie, Pharmazie, Institut für Chemie und Biochemie, AG Strukturbiochemie, Takustr. 6, 14195 Berlin, Germany Max-Planck-Institute for Biophysical Chemistry, Department of Cellular Biochemistry, Am Fassberg 11, 37077 Göttingen, Germany.
Organizational Affiliation: