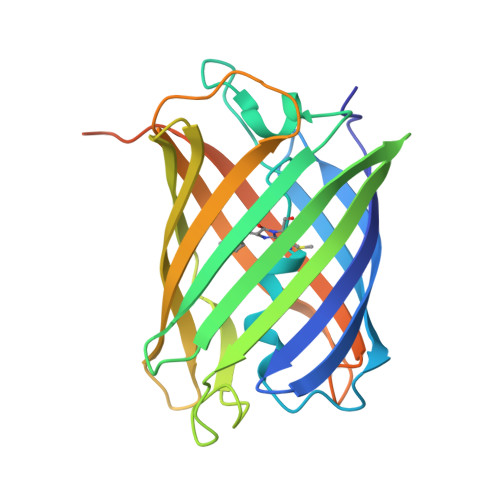
X-Ray Crystal Structure and Properties of Phanta, a Weakly Fluorescent Photochromic GFP-Like Protein.
Don Paul, C., Traore, D.A., Olsen, S., Devenish, R.J., Close, D.W., Bell, T.D., Bradbury, A., Wilce, M.C., Prescott, M.(2015) PLoS One 10: e0123338-e0123338
- PubMed: 25923520 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0123338
- Primary Citation Related Structures:
4PPJ, 4PPK, 4PPL - PubMed Abstract:
Phanta is a reversibly photoswitching chromoprotein (ΦF, 0.003), useful for pcFRET, that was isolated from a mutagenesis screen of the bright green fluorescent eCGP123 (ΦF, 0.8). We have investigated the contribution of substitutions at positions His193, Thr69 and Gln62, individually and in combination, to the optical properties of Phanta. Single amino acid substitutions at position 193 resulted in proteins with very low ΦF, indicating the importance of this position in controlling the fluorescence efficiency of the variant proteins. The substitution Thr69Val in Phanta was important for supressing the formation of a protonated chromophore species observed in some His193 substituted variants, whereas the substitution Gln62Met did not significantly contribute to the useful optical properties of Phanta. X-ray crystal structures for Phanta (2.3 Å), eCGP123T69V (2.0 Å) and eCGP123H193Q (2.2 Å) in their non-photoswitched state were determined, revealing the presence of a cis-coplanar chromophore. We conclude that changes in the hydrogen-bonding network supporting the cis-chromophore, and its contacts with the surrounding protein matrix, are responsible for the low fluorescence emission of eCGP123 variants containing a His193 substitution.
- Department of Neuro- and Sensory Physiology, University Medicine, Göttingen, 37073, Göttingen, Germany.
Organizational Affiliation: