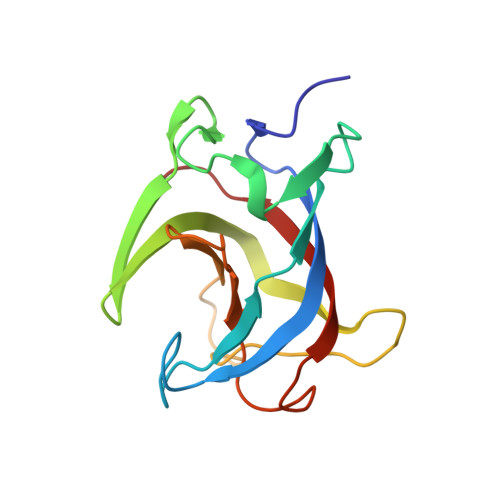
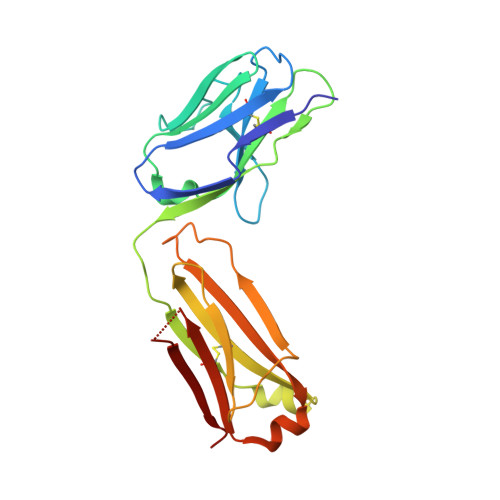
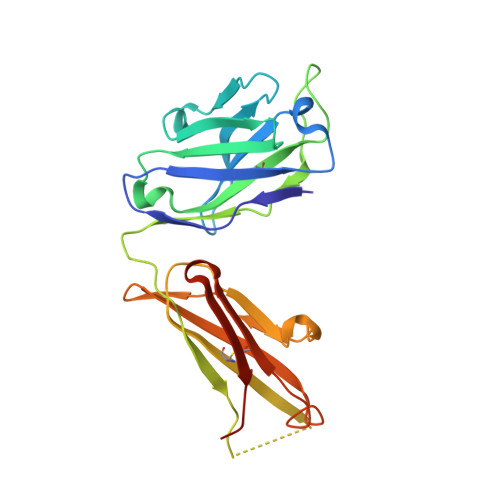
Structural basis for the neutralization of hepatitis E virus by a cross-genotype antibody
Gu, Y., Tang, X., Zhang, X., Song, C., Zheng, M., Wang, K., Zhang, J., Ng, M.H., Hew, C.L., Li, S., Xia, N., Sivaraman, J.(2015) Cell Res 25: 604-620
- PubMed: 25793314 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/cr.2015.34
- Primary Citation Related Structures:
4PLJ, 4PLK - PubMed Abstract:
Hepatitis E virus (HEV), a non-enveloped, positive-sense, single-stranded RNA virus, is a major cause of enteric hepatitis. Classified into the family Hepeviridae, HEV comprises four genotypes (genotypes 1-4), which belong to a single serotype. We describe a monoclonal antibody (mAb), 8G12, which equally recognizes all four genotypes of HEV, with ∼ 2.53-3.45 nM binding affinity. The mAb 8G12 has a protective, neutralizing capacity, which can significantly block virus infection in host cells. Animal studies with genotypes 1, 3 and 4 confirmed the cross-genotype neutralizing capacity of 8G12 and its effective prevention of hepatitis E disease. The complex crystal structures of 8G12 with the HEV E2s domain (the most protruded region of the virus capsid) of the abundant genotypes 1 and 4 were determined at 4.0 and 2.3 Å resolution, respectively. These structures revealed that 8G12 recognizes both genotypes through the epitopes in the E2s dimerization region. Structure-based mutagenesis and cell-model assays with virus-like particles identified several conserved residues (Glu549, Lys554 and Gly591) that are essential for 8G12 neutralization. Moreover, the epitope of 8G12 is identified as a key epitope involved in virus-host interactions. These findings will help develop a common strategy for the prevention of the most abundant form of HEV infection.
- 1] State Key Laboratory of Molecular Vaccinology and Molecular Diagnostics, School of Life Sciences, Xiamen University, Xiamen, Fujian 361005, China [2] National Institute of Diagnostics and Vaccine Development in Infectious Disease, School of Public Health, Xiamen University, Xiamen, Fujian 361005, China.
Organizational Affiliation: