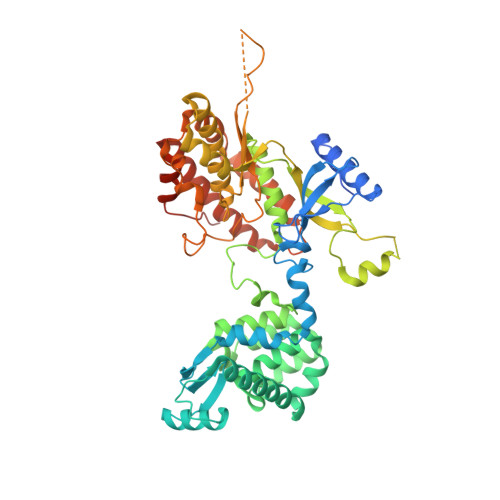
The crystal structure of the phosphatidylinositol 4-kinase II alpha.
Baumlova, A., Chalupska, D., Rozycki, B., Jovic, M., Wisniewski, E., Klima, M., Dubankova, A., Kloer, D.P., Nencka, R., Balla, T., Boura, E.(2014) EMBO Rep 15: 1085-1092
- PubMed: 25168678 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.15252/embr.201438841
- Primary Citation Related Structures:
4PLA - PubMed Abstract:
Phosphoinositides are a class of phospholipids generated by the action of phosphoinositide kinases with key regulatory functions in eukaryotic cells. Here, we present the atomic structure of phosphatidylinositol 4-kinase type IIα (PI4K IIα), in complex with ATP solved by X-ray crystallography at 2.8 Å resolution. The structure revealed a non-typical kinase fold that could be divided into N- and C-lobes with the ATP binding groove located in between. Surprisingly, a second ATP was found in a lateral hydrophobic pocket of the C-lobe. Molecular simulations and mutagenesis analysis revealed the membrane binding mode and the putative function of the hydrophobic pocket. Taken together, our results suggest a mechanism of PI4K IIα recruitment, regulation, and function at the membrane.
- Institute of Organic Chemistry and Biochemistry AS CR, Prague, Czech Republic.
Organizational Affiliation: