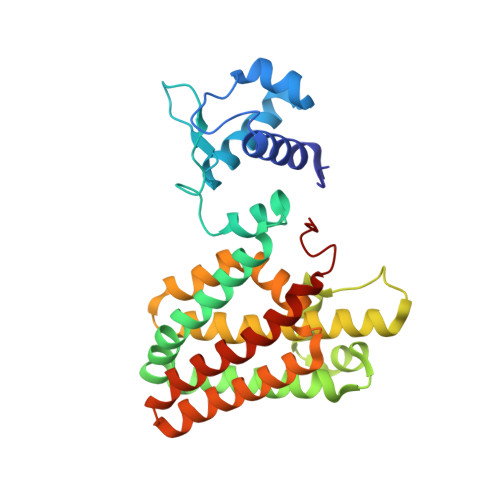
The 40-residue insertion in Vibrio cholerae FadR facilitates binding of an additional fatty acyl-CoA ligand.
Shi, W., Kovacikova, G., Lin, W., Taylor, R.K., Skorupski, K., Kull, F.J.(2015) Nat Commun 6: 6032-6032
- PubMed: 25607896 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms7032
- Primary Citation Related Structures:
4P96, 4P9U, 4PDK - PubMed Abstract:
FadR is a master regulator of fatty acid metabolism and influences virulence in certain members of Vibrionaceae. Among FadR homologues of the GntR family, the Vibrionaceae protein is unusual in that it contains a C-terminal 40-residue insertion. Here we report the structure of Vibrio cholerae FadR (VcFadR) alone, bound to DNA, and in the presence of a ligand, oleoyl-CoA. Whereas Escherichia coli FadR (EcFadR) contains only one acyl-CoA-binding site in each monomer, crystallographic and calorimetric data indicate that VcFadR has two. One of the binding sites resembles that of EcFadR, whereas the other, comprised residues from the insertion, has not previously been observed. Upon ligand binding, VcFadR undergoes a dramatic conformational change that would more fully disrupt DNA binding than EcFadR. These findings suggest that the ability to bind and respond to an additional ligand allows FadR from Vibrionaceae to function as a more efficient regulator.
- Department of Biochemistry, Geisel School of Medicine at Dartmouth, Hanover, New Hampshire 03755, USA.
Organizational Affiliation: