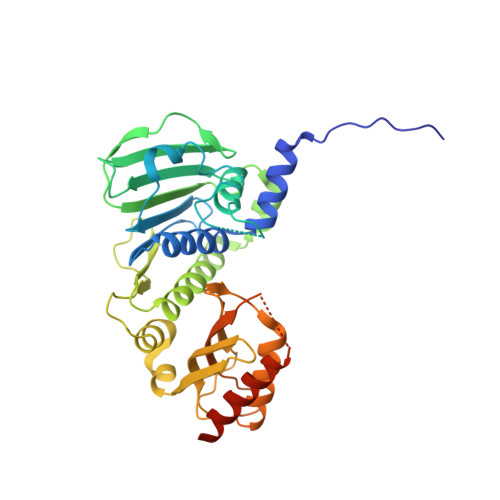
Structure of the human MLH1 N-terminus: implications for predisposition to Lynch syndrome.
Wu, H., Zeng, H., Lam, R., Tempel, W., Kerr, I.D., Min, J.(2015) Acta Crystallogr F Struct Biol Commun 71: 981-985
- PubMed: 26249686 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X15010183
- Primary Citation Related Structures:
4P7A - PubMed Abstract:
Mismatch repair prevents the accumulation of erroneous insertions/deletions and non-Watson-Crick base pairs in the genome. Pathogenic mutations in the MLH1 gene are associated with a predisposition to Lynch and Turcot's syndromes. Although genetic testing for these mutations is available, robust classification of variants requires strong clinical and functional support. Here, the first structure of the N-terminus of human MLH1, determined by X-ray crystallography, is described. The structure shares a high degree of similarity with previously determined prokaryotic MLH1 homologs; however, this structure affords a more accurate platform for the classification of MLH1 variants.
- Structural Genomics Consortium, University of Toronto, 101 College Street, Toronto, ON M5G 1L7, Canada.
Organizational Affiliation: