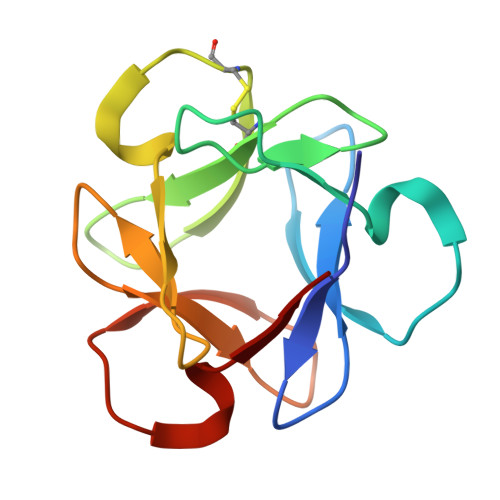
The characteristic structure of anti-HIV actinohivin in complex with three HMTG D1 chains of HIV-gp120.
Zhang, F., Hoque, M.M., Jiang, J., Suzuki, K., Tsunoda, M., Takeda, Y., Ito, Y., Kawai, G., Tanaka, H., Takenaka, A.(2014) Chembiochem 15: 2766-2773
- PubMed: 25403811 Search on PubMed
- DOI: https://doi.org/10.1002/cbic.201402352
- Primary Citation Related Structures:
4P6A - PubMed Abstract:
The anti-HIV lectin actinohivin (AH) specifically interacts with HMTG (high-mannose-type glycan), which is attached to the glycoprotein gp120 of HIV-1 in a process in which the three branched mannotriose chains (D1, D2, and D3) of HMTG exhibit different binding affinities, it being estimated that that of D1 is the strongest, that of D3 is weaker, and that of D2 is undetectable. These properties have been ascribed to the stereochemical differences in linkages between the second and the third mannose residues of the three chains. In order to clarify the interaction geometry between AH and the major target D1, an X-ray determination of the crystal structure of AH in complex with D1-which is α(1,2)mannotriose composed of three mannose (Man) residues linked together only by α(1,2) bonding-has been performed. In each of the three D1-binding pockets of AH, two Man residues of D1 are accommodated at zones 1 and 2 in the pocket, in the same way as those found in the α(1,2)mannobiose-bound AH crystals. However, an OMIT map shows poor densities at both ends of the two residues. This suggests the existence of positional disorder of D1 in the pocket: the two zones are each occupied by two Man residues in two different modes, with mode A involving the Man1 and Man2 residues and mode B the Man2 and Man3 residues. In each mode, D1 is stabilized by adopting a double-bracket-shaped conformation through CH⋅⋅⋅O interactions. In mode B, however, the Man1 residue, which is the most sensitive residue to AH binding, protrudes wholly into the solvent region without contacts with AH. In mode A, in contrast, the Man3 residue interacts with the essential hydrophobic amino acid residues (Tyr and Leu conserved between the three pockets) of AH. Therefore, mode A is likely to be the one that occurs when whole HMTG is bound. In this mode, the two hydroxy groups (O3 and O4) of the Man2 residue are anchored in zone 2 by four hydrogen bonds with Asp, Asn, and Tyr residues of AH. In addition, it has been found that an isolated water molecule buried in the hydrophobic long loop bridges between Asp of AH and the hydroxy group of Man2 through hydrogen bonds. The most interesting feature is found in the interaction of the Man1 and Man3 residues with AH. All eight hydroxy groups of the two residues are completely exposed in the solvent region, whereas their hydrophobic parts make contacts with a Leu residue and two Tyr residues so that the shape of D1 and the surface of AH fit well over a wide area. These structural characteristics are potentially useful for development of AH to produce more effective antiretroviral drugs to suppress the infectious expansion of HIV/AIDS and to help expedite an end to the HIV/AIDS pandemic in the near future.
- Graduate School of Science and Engineering, Iwaki-Meisei University, Iwaki 970-8551 (Japan).
Organizational Affiliation: