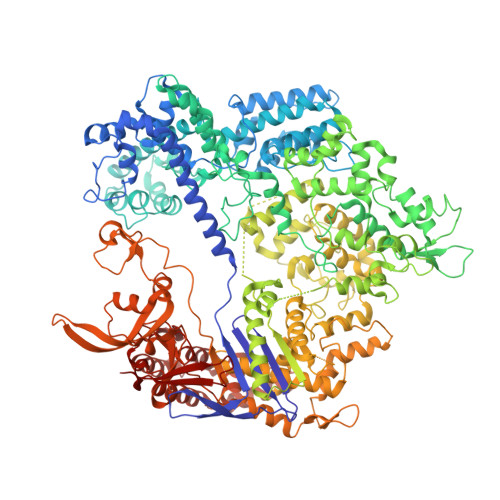
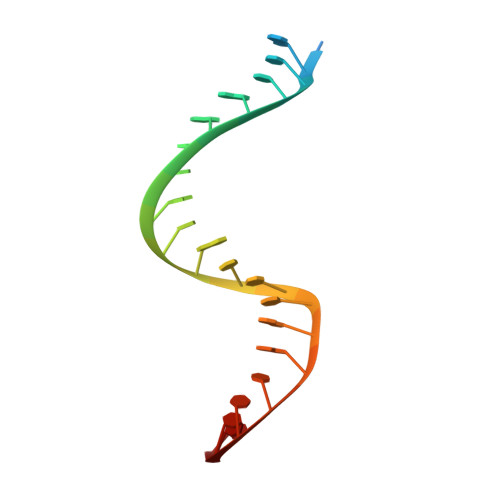
Crystal structure of Cas9 in complex with guide RNA and target DNA
Nishimasu, H., Ran, F.A., Hsu, P.D., Konermann, S., Shehata, S.I., Dohmae, N., Ishitani, R., Zhang, F., Nureki, O.(2014) Cell 156: 935-949
- PubMed: 24529477 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.cell.2014.02.001
- Primary Citation Related Structures:
4OO8 - PubMed Abstract:
The CRISPR-associated endonuclease Cas9 can be targeted to specific genomic loci by single guide RNAs (sgRNAs). Here, we report the crystal structure of Streptococcus pyogenes Cas9 in complex with sgRNA and its target DNA at 2.5 Å resolution. The structure revealed a bilobed architecture composed of target recognition and nuclease lobes, accommodating the sgRNA:DNA heteroduplex in a positively charged groove at their interface. Whereas the recognition lobe is essential for binding sgRNA and DNA, the nuclease lobe contains the HNH and RuvC nuclease domains, which are properly positioned for cleavage of the complementary and noncomplementary strands of the target DNA, respectively. The nuclease lobe also contains a carboxyl-terminal domain responsible for the interaction with the protospacer adjacent motif (PAM). This high-resolution structure and accompanying functional analyses have revealed the molecular mechanism of RNA-guided DNA targeting by Cas9, thus paving the way for the rational design of new, versatile genome-editing technologies.
- Department of Biophysics and Biochemistry, Graduate School of Science, The University of Tokyo, 2-11-16 Yayoi, Bunkyo, Tokyo 113-0032, Japan; JST, PRESTO, 2-11-16 Yayoi, Bunkyo, Tokyo 113-0032, Japan.
Organizational Affiliation: