Structural and Thermodynamic Characterization of Cadherin beta-Catenin alpha-Catenin Complex Formation.
Pokutta, S., Choi, H.J., Ahlsen, G., Hansen, S.D., Weis, W.I.(2014) J Biological Chem 289: 13589-13601
- PubMed: 24692547 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M114.554709
- Primary Citation Related Structures:
4ONS - PubMed Abstract:
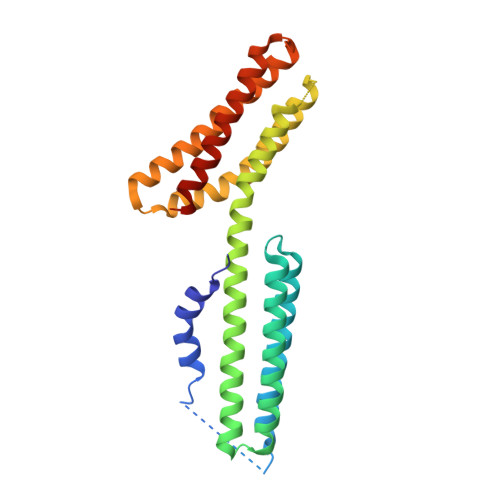
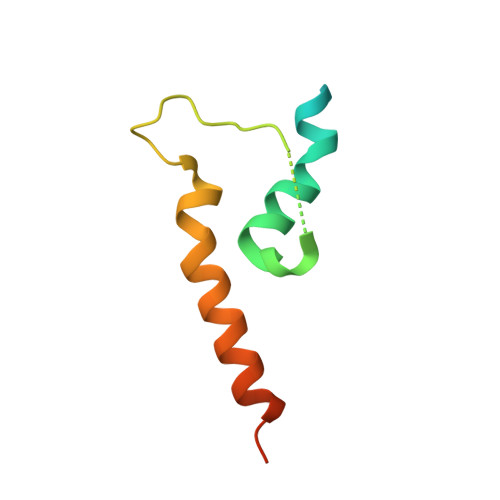
The classical cadherin·β-catenin·α-catenin complex mediates homophilic cell-cell adhesion and mechanically couples the actin cytoskeletons of adjacent cells. Although α-catenin binds to β-catenin and to F-actin, β-catenin significantly weakens the affinity of α-catenin for F-actin. Moreover, α-catenin self-associates into homodimers that block β-catenin binding. We investigated quantitatively and structurally αE- and αN-catenin dimer formation, their interaction with β-catenin and the cadherin·β-catenin complex, and the effect of the α-catenin actin-binding domain on β-catenin association. The two α-catenin variants differ in their self-association properties: at physiological temperatures, αE-catenin homodimerizes 10× more weakly than does αN-catenin but is kinetically trapped in its oligomeric state. Both αE- and αN-catenin bind to β-catenin with a Kd of 20 nM, and this affinity is increased by an order of magnitude when cadherin is bound to β-catenin. We describe the crystal structure of a complex representing the full β-catenin·αN-catenin interface. A three-dimensional model of the cadherin·β-catenin·α-catenin complex based on these new structural data suggests mechanisms for the enhanced stability of the ternary complex. The C-terminal actin-binding domain of α-catenin has no influence on the interactions with β-catenin, arguing against models in which β-catenin weakens actin binding by stabilizing inhibitory intramolecular interactions between the actin-binding domain and the rest of α-catenin.
- From the Departments of Structural Biology and Molecular and Cellular Physiology, Stanford University, Stanford, California 94305.
Organizational Affiliation: