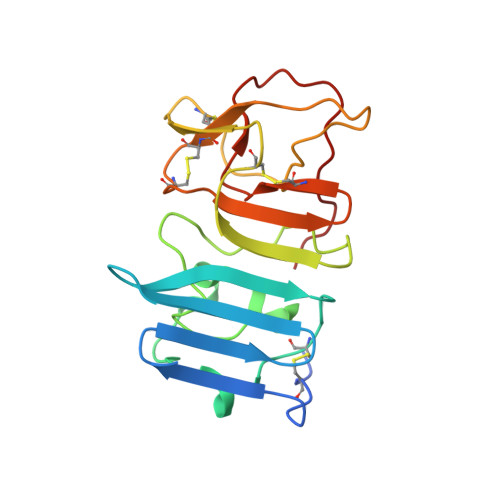
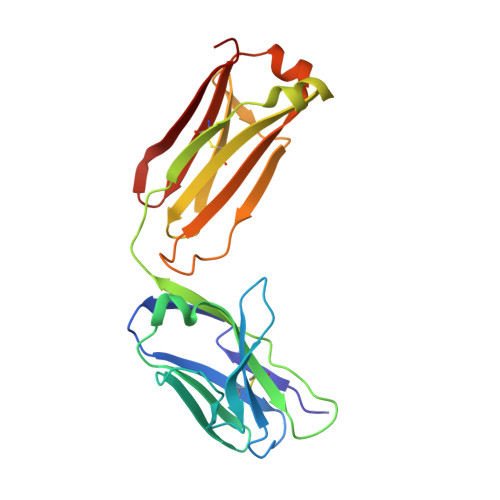
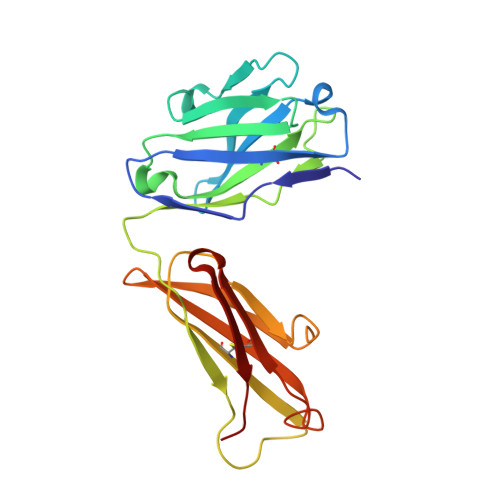
Structural basis of Flavivirus NS1 assembly and antibody recognition.
Edeling, M.A., Diamond, M.S., Fremont, D.H.(2014) Proc Natl Acad Sci U S A 111: 4285-4290
- PubMed: 24594604 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1322036111
- Primary Citation Related Structures:
4OIE, 4OIG, 4OII - PubMed Abstract:
The Flavivirus nonstructural protein 1 (NS1) is a conserved, membrane-associated and secreted glycoprotein with replication and immune evasion functions. Secreted NS1 is a hexameric, barrel-shaped lipoprotein that can bind back to the plasma membrane of cells. Antibodies targeting cell surface-associated NS1 can be protective in vivo in a manner dependent on Fc effector functions. We describe here the crystal structure of a C-terminal fragment (residues 172-352) of West Nile (WNV) and Dengue virus NS1 proteins at 1.85 and 2.7 Å resolution, respectively. NS1(172-352) assembles as a unique rod-shaped dimer composed of a 16-stranded β-platform flanked on one face by protruding connecting loops. We also determined the 3.0 Å resolution structure of WNV NS1(172-352) with the protective 22NS1 antibody Fab, which engages the loop-face of the rod. The head-to-head NS1(172-352) dimer we observe in crystal lattices is supported by multiangle light and small-angle X-ray scattering studies. We used the available cryo-electron microscopy reconstruction to develop a pseudoatomic model of the NS1 hexamer. The model was constructed with the NS1(172-352) dimeric rod aligned with the long axis of the barrel, and with the loop-face oriented away from the core. Difference densities suggest that the N-terminal region of NS1 forms globular lobes that mediate lateral contacts between dimers in the hexamer. Our model also suggests that the N-terminal lobe forms the surface of the central cavity where lipid binding may occur.
- Departments of Pathology and Immunology, Medicine, Molecular Microbiology, and Biochemistry and Molecular Biophysics, Washington University in St. Louis, St. Louis, MO 63110.
Organizational Affiliation: