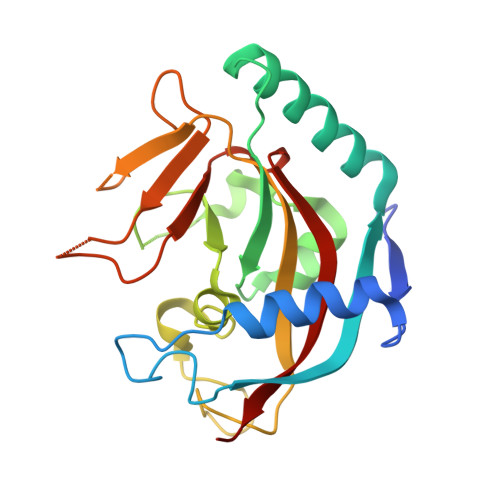
Disruption of Wnt/ beta-Catenin Signaling and Telomeric Shortening Are Inextricable Consequences of Tankyrase Inhibition in Human Cells.
Kulak, O., Chen, H., Holohan, B., Wu, X., He, H., Borek, D., Otwinowski, Z., Yamaguchi, K., Garofalo, L.A., Ma, Z., Wright, W., Chen, C., Shay, J.W., Zhang, X., Lum, L.(2015) Mol Cell Biol 35: 2425-2435
- PubMed: 25939383 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/MCB.00392-15
- Primary Citation Related Structures:
4OA7, 4TOR, 4TOS - PubMed Abstract:
Maintenance of chromosomal ends (telomeres) directly contributes to cancer cell immortalization. The telomere protection enzymes belonging to the tankyrase (Tnks) subfamily of poly(ADP-ribose) polymerases (PARPs) have recently been shown to also control transcriptional response to secreted Wnt signaling molecules. Whereas Tnks inhibitors are currently being developed as therapeutic agents for targeting Wnt-related cancers and as modulators of Wnt signaling in tissue-engineering agendas, their impact on telomere length maintenance remains unclear. Here, we leveraged a collection of Wnt pathway inhibitors with previously unassigned mechanisms of action to identify novel pharmacophores supporting Tnks inhibition. A multifaceted experimental approach that included structural, biochemical, and cell biological analyses revealed two distinct chemotypes with selectivity for Tnks enzymes. Using these reagents, we revealed that Tnks inhibition rapidly induces DNA damage at telomeres and telomeric shortening upon long-term chemical exposure in cultured cells. On the other hand, inhibitors of the Wnt acyltransferase Porcupine (Porcn) elicited neither effect. Thus, Tnks inhibitors impact telomere length maintenance independently of their affects on Wnt/β-catenin signaling. We discuss the implications of these findings for anticancer and regenerative medicine agendas dependent upon chemical inhibitors of Wnt/β-catenin signaling.
- Department of Cell Biology, University of Texas Southwestern Medical Center, Dallas, Texas, USA.
Organizational Affiliation: