Structure of CHD4 double chromodomains depicts cooperative folding for DNA binding
Wiggs, K.R., Chruszcz, M., Su, X., Minor, W., Khorasanizadeh, S.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Chromodomain-helicase-DNA-binding protein 4 | A [auth X] | 202 | Homo sapiens | Mutation(s): 0 Gene Names: CHD4 EC: 3.6.4.12 (PDB Primary Data), 3.6.4 (UniProt) | 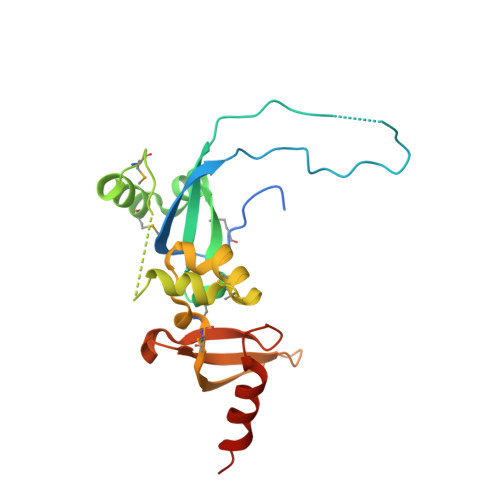 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q14839 GTEx: ENSG00000111642 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q14839 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Modified Residues 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| MSE Query on MSE | A [auth X] | L-PEPTIDE LINKING | C5 H11 N O2 Se |  | MET |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 50.211 | α = 90 |
| b = 60.475 | β = 90 |
| c = 92.988 | γ = 90 |
| Software Name | Purpose |
|---|---|
| MAR345 | data collection |
| SERGUI | data collection |
| HKL-3000 | phasing |
| SHELXCD | phasing |
| SHELXE | model building |
| MLPHARE | phasing |
| DM | model building |
| RESOLVE | model building |
| ARP/wARP | model building |
| CCP4 | model building |
| REFMAC | refinement |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| DM | phasing |
| RESOLVE | phasing |
| CCP4 | phasing |