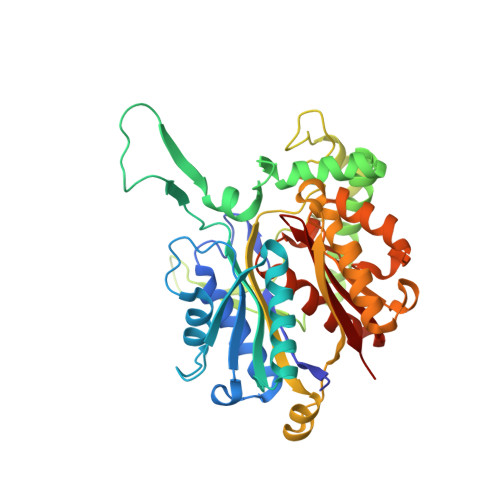
Crystal structure and biochemical characterization of PhaA from Ralstonia eutropha, a polyhydroxyalkanoate-producing bacterium.
Kim, E.J., Kim, K.J.(2014) Biochem Biophys Res Commun 452: 124-129
- PubMed: 25152395 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2014.08.074
- Primary Citation Related Structures:
4O99, 4O9A, 4O9C - PubMed Abstract:
PhaA from Ralstonia eutropha (RePhaA) is the first enzyme in the polyhydroxyalbutyrate (PHB) biosynthetic pathway and catalyzes the condensation of two molecules of acetyl-CoA to acetoacetyl-CoA. To investigate the molecular mechanism underlying PHB biosynthesis, we determined the crystal structures of the RePhaA protein in apo- and CoA-bound forms. The RePhaA structure adopts the type II biosynthetic thiolase fold forming a tetramer by means of dimerization of two dimers. The crystal structure of RePhaA in complex with CoA revealed that the enzyme contained a unique Phe219 residue, resulting that the ADP moiety binds in somewhat different position compared with that bound in other thiolase enzymes. Our study provides structural insight into the substrate specificity of RePhaA. Results indicate the presence of a small pocket near the Cys88 covalent catalytic residue leading to the possibility of the enzyme to accommodate acetyl-CoA as a sole substrate instead of larger acyl-CoA molecules such as propionyl-CoA. Furthermore, the roles of key residues involved in substrate binding and enzyme catalysis were confirmed by site-directed mutagenesis.
- School of Life Sciences, KNU Creative BioResearch Group, Kyungpook National University, Daehak-ro 80, Buk-ku, Daegu 702-701, Republic of Korea.
Organizational Affiliation: