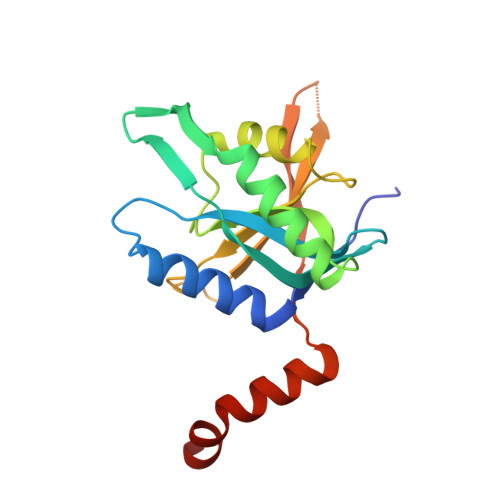
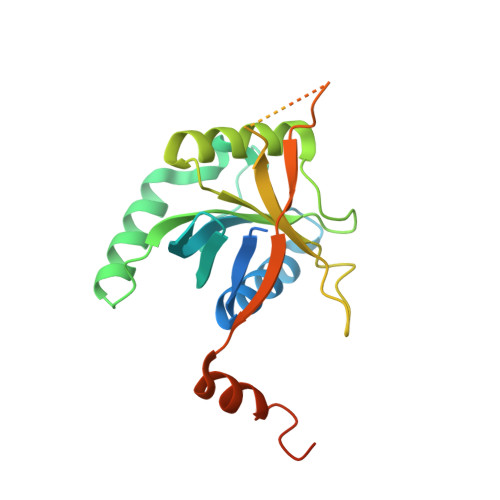
Structure of the Rpn11-Rpn8 dimer reveals mechanisms of substrate deubiquitination during proteasomal degradation.
Worden, E.J., Padovani, C., Martin, A.(2014) Nat Struct Mol Biol 21: 220-227
- PubMed: 24463465 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.2771
- Primary Citation Related Structures:
4O8X, 4O8Y - PubMed Abstract:
Polyubiquitin chains target protein substrates to the 26S proteasome, where they are removed by the deubiquitinase Rpn11 to allow efficient substrate degradation. Despite Rpn11's essential function during substrate processing, its detailed structural and biochemical characterization has been hindered by difficulties in purifying the isolated enzyme. Here we report the 2.0-Å crystal structures of Zn(2+)-free and Zn(2+)-bound Saccharomyces cerevisiae Rpn11 in an MPN-domain heterodimer with Rpn8. The Rpn11-Rpn8 interaction occurs via two distinct interfaces that may be conserved in related MPN-domain complexes. Our structural and mutational studies reveal that Rpn11 lacks a conserved surface to bind the ubiquitin Ile44 patch, does not interact with the moiety on the proximal side of the scissile isopeptide bond and exhibits no linkage specificity for ubiquitin cleavage. These findings explain how Rpn11 functions as a promiscuous deubiquitinase for cotranslocational substrate deubiquitination during proteasomal degradation.
- Department of Molecular and Cell Biology, University of California, Berkeley, Berkeley, California, USA.
Organizational Affiliation: