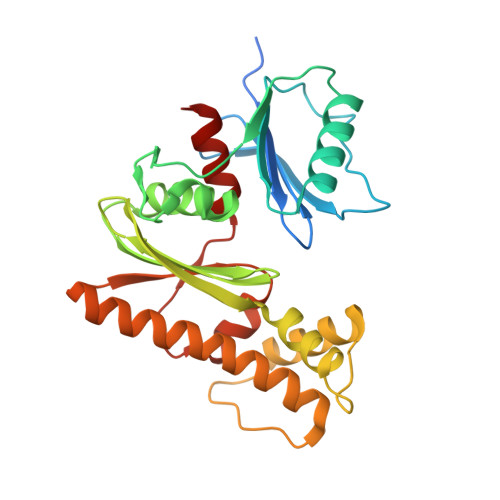
Crystal structure of Type III pantothenate kinase from Burkholderia thailandensis in complex with pantothenate and phosphate
Abendroth, J., Dranow, D.M., Lorimer, D., Fairman, J.W., Edwards, T.E., Seattle Structural Genomics Center for Infectious Disease (SSGCID)To be published.