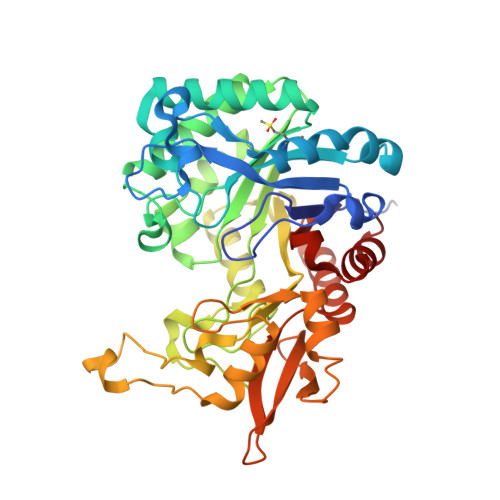
Structure of chitinase D from Serratia proteamaculans reveals the structural basis of its dual action of hydrolysis and transglycosylation
Madhuprakash, J., Singh, A., Kumar, S., Sinha, M., Kaur, P., Sharma, S., Podile, A.R., Singh, T.P.(2013) Int J Biochem Mol Biol 4: 166-178
- PubMed: 24380021 Search on PubMedSearch on PubMed Central
- Primary Citation Related Structures:
4NZC - PubMed Abstract:
Chitinases are known to hydrolyze chitin polymers into smaller chitooligosaccharides. Chitinase from bacterium Serratia proteamaculans (SpChiD) is found to exhibit both hydrolysis and transglycosylation activities. SpChiD belongs to family 18 of glycosyl hydrolases (GH-18). The recombinant SpChiD was crystallized and its three-dimensional structure was determined at 1.49 Å resolution. The structure was refined to an R-factor of 16.2%. SpChiD consists of 406 amino acid residues. The polypeptide chain of SpChiD adopts a (β/α)8 triosephosphate isomerase (TIM) barrel structure. SpChiD contains three acidic residues, Asp149, Asp151 and Glu153 as part of its catalytic scheme. While both Asp149 and Glu153 adopt single conformations, Asp151 is observed in two conformations. The substrate binding cleft is partially obstructed by a protruding loop, Asn30 - Asp42 causing a considerable reduction in the number of available subsites in the substrate binding site. The positioning of loop, Asn30 - Asp42 appears to be responsible for the transglycosylation activity. The structure determination indicated the presence of sulfone Met89 (SMet89). The sulfone methionine residue is located on the surface of the protein at a site where extra domain is attached in other chitinases. This is the first structure of a single domain chitinase with hydrolytic and transglycosylation activities.
- Department of Plant Sciences, School of Life Sciences, University of Hyderabad Hyderabad, India.
Organizational Affiliation: