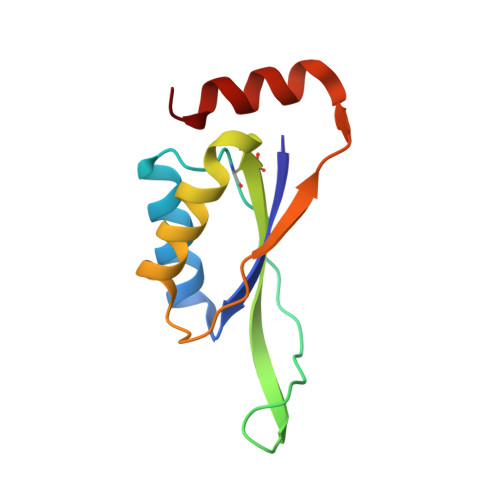
The structures of the CutA1 proteins from Thermus thermophilus and Pyrococcus horikoshii: characterization of metal-binding sites and metal-induced assembly
Bagautdinov, B.(2014) Acta Crystallogr F Struct Biol Commun 70: 404-413
- PubMed: 24699729 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X14003422
- Primary Citation Related Structures:
1NZA, 1V6H, 4NYO - PubMed Abstract:
CutA1 (copper tolerance A1) is a widespread cytoplasmic protein found in archaea, bacteria, plants and animals, including humans. In Escherichia coli it is implicated in divalent metal tolerance, while the mammalian CutA1 homologue has been proposed to mediate brain enzyme acetylcholinesterase activity and copper homeostasis. The X-ray structures of CutA1 from the thermophilic bacterium Thermus thermophilus (TtCutA1) with and without bound Na(+) at 1.7 and 1.9 Å resolution, respectively, and from the hyperthermophilic archaeon Pyrococcus horikoshii (PhCutA1) in complex with Na(+) at 1.8 Å resolution have been determined. Both are short and rigid proteins of about 12 kDa that form intertwined compact trimers in the crystal and solution. The main difference in the structures is a wide-type β-bulge on top of the TtCutA1 trimer. It affords a mechanism for lodging a single-residue insertion in the middle of β2 while preserving the interprotomer main-chain hydrogen-bonding network. The liganded forms of the proteins provide new structural information about the metal-binding sites and CutA1 assembly. The Na(+)-TtCutA1 structure unveils a dodecameric assembly with metal ions in the trimer-trimer interfaces and the lateral clefts of the trimer. For Na(+)-PhCutA1, the metal ion associated with six waters in an octahedral geometry. The structures suggest that CutA1 may contribute to regulating intracellular metal homeostasis through various binding modes.
- Japan Synchrotron Radiation Research Institute (JASRI/SPring-8), 1-1-1 Kouto, Sayo, Hyogo 679-5198, Japan.
Organizational Affiliation: