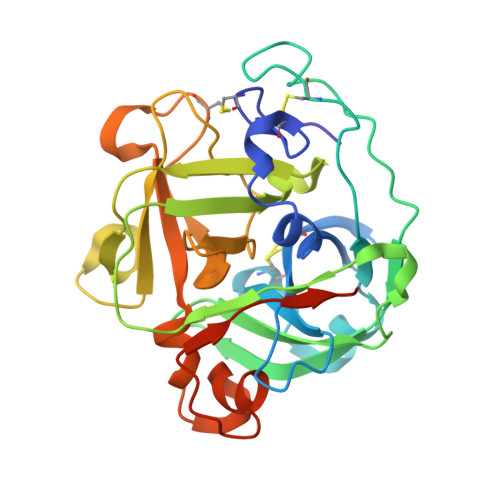
Atomic resolution structure of a lysine-specific endoproteinase from Lysobacter enzymogenes suggests a hydroxyl group bound to the oxyanion hole.
Asztalos, P., Muller, A., Holke, W., Sobek, H., Rudolph, M.G.(2014) Acta Crystallogr D Biol Crystallogr 70: 1832-1843
- PubMed: 25004961 Search on PubMed
- DOI: https://doi.org/10.1107/S1399004714008463
- Primary Citation Related Structures:
4NSV, 4NSY - PubMed Abstract:
Lysobacter enzymogenes lysyl endoproteinase (LysC) is a trypsin-type serine protease with a high pH optimum that hydrolyses all Lys-Xaa peptide bonds. The high specificity of LysC renders it useful for biotechnological purposes. The K30R variant of a related lysyl endoproteinase from Achromobacter lyticus has favourable enzymatic properties that might be transferrable to LysC. To visualize structural differences in the substrate-binding sites, the crystal structures of wild-type and the K30R variant of LysC were determined. The mutation is located at a distance of 12 Å from the catalytic triad and subtly changes the surface properties of the substrate-binding site. The high pH optimum of LysC can be attributed to electrostatic effects of an aromatic Tyr/His stack on the catalytic aspartate and is a general feature of this enzyme subfamily. LysC crystals in complex with the covalent inhibitor N(α)-p-tosyl-lysyl chloromethylketone yielded data to 1.1 and 0.9 Å resolution, resulting in unprecedented precision of the active and substrate-binding sites for this enzyme subfamily. Error estimates on bond lengths and difference electron density indicate that instead of the expected oxyanion a hydroxyl group binds to the partially solvent-exposed oxyanion hole. Protonation of the alkoxide catalytic intermediate might be a recurring feature during serine protease catalysis.
- Roche Diagnostics GmbH, Nonnenwald 2, 82377 Penzberg, Germany.
Organizational Affiliation: