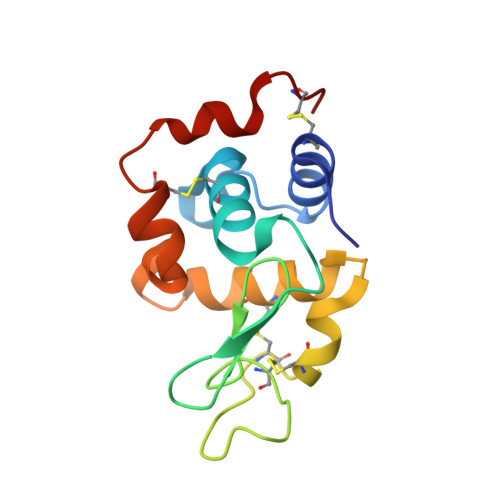
Carboplatin binding to histidine.
Tanley, S.W., Diederichs, K., Kroon-Batenburg, L.M., Levy, C., Schreurs, A.M., Helliwell, J.R.(2014) Acta Crystallogr F Struct Biol Commun 70: 1135-1142
- PubMed: 25195881 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X14016161
- Primary Citation Related Structures:
4LT0, 4LT3, 4NSG, 4NSH, 4NSI, 4NSJ - PubMed Abstract:
Carboplatin is a second-generation platinum anticancer agent used for the treatment of a variety of cancers. Previous X-ray crystallographic studies of carboplatin binding to histidine (in hen egg-white lysozyme; HEWL) showed the partial conversion of carboplatin to cisplatin owing to the high NaCl concentration used in the crystallization conditions. HEWL co-crystallizations with carboplatin in NaBr conditions have now been carried out to confirm whether carboplatin converts to the bromine form and whether this takes place in a similar way to the partial conversion of carboplatin to cisplatin observed previously in NaCl conditions. Here, it is reported that a partial chemical transformation takes place but to a transplatin form. Thus, to attempt to resolve purely carboplatin binding at histidine, this study utilized co-crystallization of HEWL with carboplatin without NaCl to eliminate the partial chemical conversion of carboplatin. Tetragonal HEWL crystals co-crystallized with carboplatin were successfully obtained in four different conditions, each at a different pH value. The structural results obtained show carboplatin bound to either one or both of the N atoms of His15 of HEWL, and this particular variation was dependent on the concentration of anions in the crystallization mixture and the elapsed time, as well as the pH used. The structural details of the bound carboplatin molecule also differed between them. Overall, the most detailed crystal structure showed the majority of the carboplatin atoms bound to the platinum centre; however, the four-carbon ring structure of the cyclobutanedicarboxylate moiety (CBDC) remained elusive. The potential impact of the results for the administration of carboplatin as an anticancer agent are described.
- School of Chemistry, Faculty of Engineering and Physical Sciences, University of Manchester, Brunswick Street, Manchester M13 9PL, England.
Organizational Affiliation: