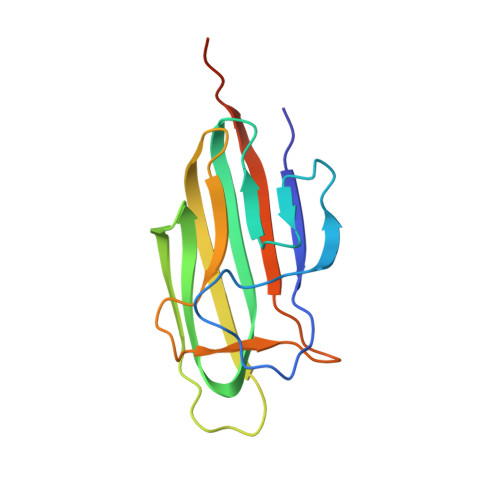
The macular degeneration-linked C1QTNF5 (S163) mutation causes higher-order structural rearrangements.
Tu, X., Palczewski, K.(2014) J Struct Biol 186: 86-94
- PubMed: 24531000 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jsb.2014.02.001
- Primary Citation Related Structures:
4NN0 - PubMed Abstract:
The C1q-tumor necrosis factor 5 (C1QTNF5) protein plays a significant role in retinal pigmented epithelium (RPE) cellular adhesion. The C1QTNF5 gene is co-transcribed with the frizzled-related protein (MFRP) gene. A Ser-to-Arg mutation at site 163 (S163R) in C1QTNF5 is known to cause late-onset retinal macular degeneration (L-ORMD). Here we also found that C1QTNF5 monomers can multimerize into a bouquet-like octadecamer. We found that a novel intermolecular hydrogen-bond network of S163 that glues adjacent globular heads of C1QTNF5 together was weakened or abolished by the R163 pathogenic mutation. These findings could underlie the structural basis of this protein's adhesive function and relate to the pathogenesis of its S163R mutation. Additionally, the fact that C1QTNF5 immobilized to a resin selectively enriched detergent extracted membrane-bound MFRP, further confirmed their interaction, implying functions other than cellular adhesion for C1QTNF5.
- Department of Pharmacology, School of Medicine, Case Western Reserve University, USA.
Organizational Affiliation: