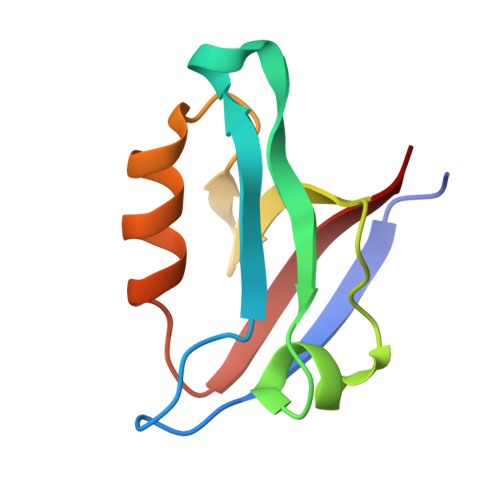
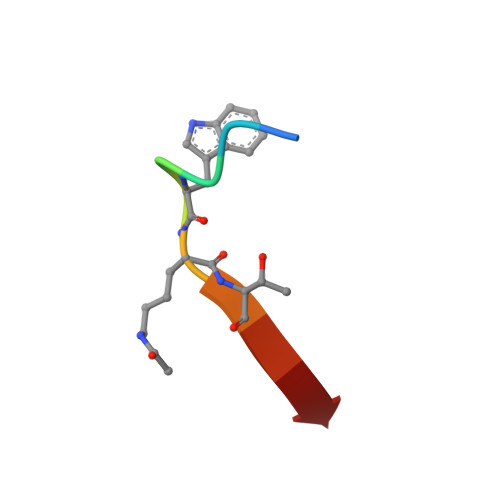
Chemically Modified Peptide Scaffolds Target the CFTR-Associated Ligand PDZ Domain.
Amacher, J.F., Zhao, R., Spaller, M.R., Madden, D.R.(2014) PLoS One 9: e103650-e103650
- PubMed: 25136860 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0103650
- Primary Citation Related Structures:
4NMO, 4NMP, 4NMQ, 4NMR, 4NMS, 4NMT, 4NMV - PubMed Abstract:
PDZ domains are protein-protein interaction modules that coordinate multiple signaling and trafficking pathways in the cell and that include active therapeutic targets for diseases such as cancer, cystic fibrosis, and addiction. Our previous work characterized a PDZ interaction that restricts the apical membrane half-life of the cystic fibrosis transmembrane conductance regulator (CFTR). Using iterative cycles of peptide-array and solution-binding analysis, we targeted the PDZ domain of the CFTR-Associated Ligand (CAL), and showed that an engineered peptide inhibitor rescues cell-surface expression of the most common CFTR disease mutation ΔF508. Here, we present a series of scaffolds containing chemically modifiable side chains at all non-motif positions along the CAL PDZ domain binding cleft. Concordant equilibrium dissociation constants were determined in parallel by fluorescence polarization, isothermal titration calorimetry, and surface plasmon resonance techniques, confirming robust affinity for each scaffold and revealing an enthalpically driven mode of inhibitor binding. Structural studies demonstrate a conserved binding mode for each peptide, opening the possibility of combinatorial modification. Finally, we diversified one of our peptide scaffolds with halogenated substituents that yielded modest increases in binding affinity. Overall, this work validates our approach and provides a stereochemical foundation for further CAL inhibitor design and screening.
- Department of Biochemistry, Geisel School of Medicine at Dartmouth, Hanover, New Hampshire, United States of America.
Organizational Affiliation: