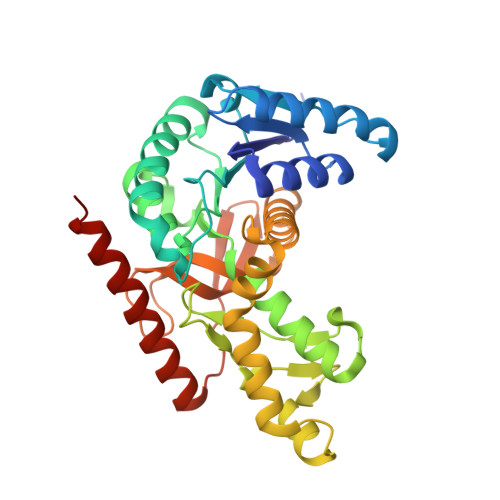
Biochemical and structural characterization of Cryptosporidium parvum Lactate dehydrogenase.
Cook, W.J., Senkovich, O., Hernandez, A., Speed, H., Chattopadhyay, D.(2014) Int J Biol Macromol 74C: 608-619
- PubMed: 25542170 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2014.12.019
- Primary Citation Related Structures:
4ND1, 4ND2, 4ND3, 4ND4, 4ND5 - PubMed Abstract:
The protozoan parasite Cryptosporidium parvum causes waterborne diseases worldwide. There is no effective therapy for C. parvum infection. The parasite depends mainly on glycolysis for energy production. Lactate dehydrogenase is a major regulator of glycolysis. This paper describes the biochemical characterization of C. parvum lactate dehydrogenase and high resolution crystal structures of the apo-enzyme and four ternary complexes. The ternary complexes capture the enzyme bound to NAD/NADH or its 3-acetylpyridine analog in the cofactor binding pocket, while the substrate binding site is occupied by one of the following ligands: lactate, pyruvate or oxamate. The results reveal distinctive features of the parasitic enzyme. For example, C. parvum lactate dehydrogenase prefers the acetylpyridine analog of NADH as a cofactor. Moreover, it is slightly less sensitive to gossypol inhibition compared with mammalian lactate dehydrogenases and not inhibited by excess pyruvate. The active site loop and the antigenic loop in C. parvum lactate dehydrogenase are considerably different from those in the human counterpart. Structural features and enzymatic properties of C. parvum lactate dehydrogenase are similar to enzymes from related parasites. Structural comparison with malate dehydrogenase supports a common ancestry for the two genes.
- Department of Pathology, University of Alabama at Birmingham, Birmingham, AL 35294, United States.
Organizational Affiliation: