Towards a new combination therapy for tuberculosis with next generation benzothiazinones.
Makarov, V., Lechartier, B., Zhang, M., Neres, J., van der Sar, A.M., Raadsen, S.A., Hartkoorn, R.C., Ryabova, O.B., Vocat, A., Decosterd, L.A., Widmer, N., Buclin, T., Bitter, W., Andries, K., Pojer, F., Dyson, P.J., Cole, S.T.(2014) EMBO Mol Med 6: 372-383
- PubMed: 24500695 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/emmm.201303575
- Primary Citation Related Structures:
4NCR - PubMed Abstract:
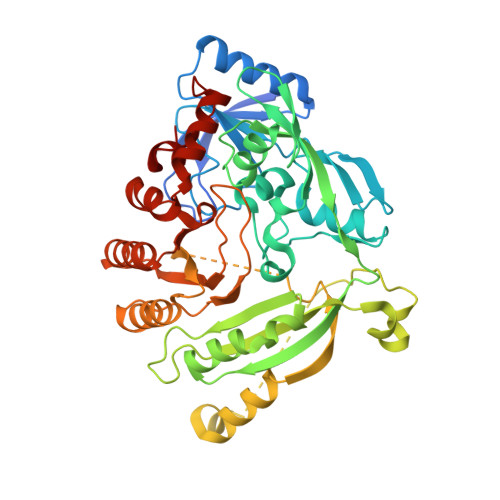
The benzothiazinone lead compound, BTZ043, kills Mycobacterium tuberculosis by inhibiting the essential flavo-enzyme DprE1, decaprenylphosphoryl-beta-D-ribose 2-epimerase. Here, we synthesized a new series of piperazine-containing benzothiazinones (PBTZ) and show that, like BTZ043, the preclinical candidate PBTZ169 binds covalently to DprE1. The crystal structure of the DprE1-PBTZ169 complex reveals formation of a semimercaptal adduct with Cys387 in the active site and explains the irreversible inactivation of the enzyme. Compared to BTZ043, PBTZ169 has improved potency, safety and efficacy in zebrafish and mouse models of tuberculosis (TB). When combined with other TB drugs, PBTZ169 showed additive activity against M. tuberculosis in vitro except with bedaquiline (BDQ) where synergy was observed. A new regimen comprising PBTZ169, BDQ and pyrazinamide was found to be more efficacious than the standard three drug treatment in a murine model of chronic disease. PBTZ169 is thus an attractive drug candidate to treat TB in humans.
- More Medicines for Tuberculosis (MM4TB) Consortium www.mm4tb.org.
Organizational Affiliation: