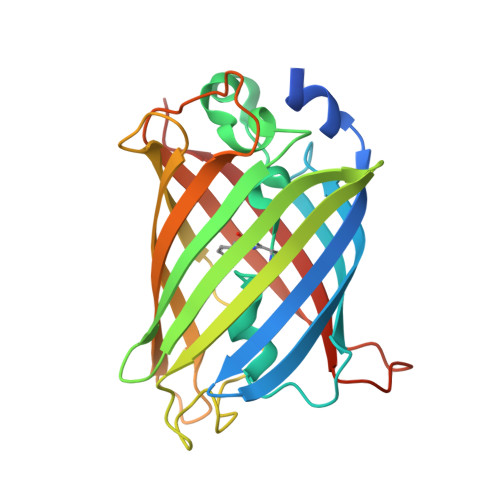
Three dimensional structure of the dimeric gene-engineered variant of green fluorescent protein egfp-K162Q in P61 crystal space group
Pletneva, N.V., V Pletnev, S., Bogdanov, A.M., Goryacheva, E.A., Artemyev, I.V., Souslova, E.A., Arkhipova, S.F., Pletnev, V.Z.(2014) Rus J Bioorg Chem 40: 383-389