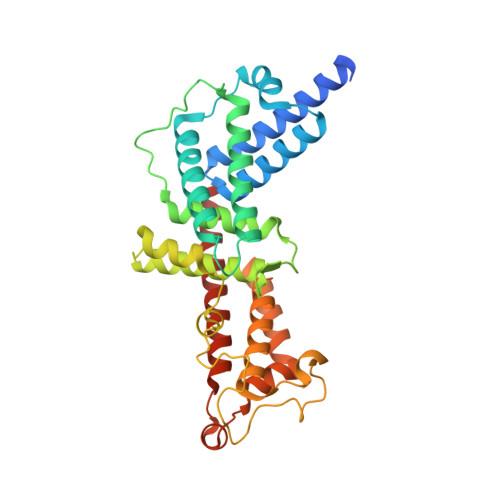
Structural Basis for PI(4)P-Specific Membrane Recruitment of the Legionella pneumophila Effector DrrA/SidM.
Del Campo, C.M., Mishra, A.K., Wang, Y.H., Roy, C.R., Janmey, P.A., Lambright, D.G.(2014) Structure 22: 397-408
- PubMed: 24530282 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2013.12.018
- Primary Citation Related Structures:
4MXP - PubMed Abstract:
Recruitment of the Legionella pneumophila effector DrrA to the Legionella-containing vacuole, where it activates and AMPylates Rab1, is mediated by a P4M domain that binds phosphatidylinositol 4-phosphate [PI(4)P] with high affinity and specificity. Despite the importance of PI(4)P in Golgi trafficking and its manipulation by pathogens, the structural bases for PI(4)P-dependent membrane recruitment remain poorly defined. Here, we determined the crystal structure of a DrrA fragment including the P4M domain in complex with dibutyl PI(4)P and investigated the determinants of phosphoinositide recognition and membrane targeting. Headgroup recognition involves an elaborate network of direct and water-mediated interactions with basic and polar residues in the context of a deep, constrictive binding pocket. An adjacent hydrophobic helical element packs against the acyl chains and inserts robustly into PI(4)P-containing monolayers. The structural, biochemical, and biophysical data reported here support a detailed structural mechanism for PI(4)P-dependent membrane targeting by DrrA.
- Program in Molecular Medicine and Department of Biochemistry and Molecular Pharmacology, University of Massachusetts Medical School, Worcester, MA 01605, USA.
Organizational Affiliation: