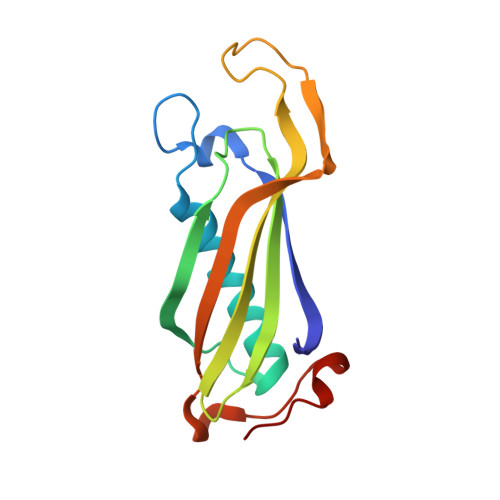
High resolution crystal structure of Clostridium propionicum beta-alanyl-CoA:ammonia lyase, a new member of the "hot dog fold" protein superfamily.
Heine, A., Herrmann, G., Selmer, T., Terwesten, F., Buckel, W., Reuter, K.(2014) Proteins 82: 2041-2053
- PubMed: 24623648 Search on PubMed
- DOI: https://doi.org/10.1002/prot.24557
- Primary Citation Related Structures:
4MTU, 4MZQ - PubMed Abstract:
Clostridium propionicum is the only organism known to ferment β-alanine, a constituent of coenzyme A (CoA) and the phosphopantetheinyl prosthetic group of holo-acyl carrier protein. The first step in the fermentation is a CoA-transfer to β-alanine. Subsequently, the resulting β-alanyl-CoA is deaminated by the enzyme β-alanyl-CoA:ammonia lyase (Acl) to reversibly form ammonia and acrylyl-CoA. We have determined the crystal structure of Acl in its apo-form at a resolution of 0.97 Å as well as in complex with CoA at a resolution of 1.59 Å. The structures reveal that the enyzme belongs to a superfamily of proteins exhibiting a so called "hot dog fold" which is characterized by a five-stranded antiparallel β-sheet with a long α-helix packed against it. The functional unit of all "hot dog fold" proteins is a homodimer containing two equivalent substrate binding sites which are established by the dimer interface. In the case of Acl, three functional dimers combine to a homohexamer strongly resembling the homohexamer formed by YciA-like acyl-CoA thioesterases. Here, we propose an enzymatic mechanism based on the crystal structure of the Acl·CoA complex and molecular docking.
- Institut für Pharmazeutische Chemie, Fachbereich Pharmazie, Philipps-Universität Marburg, Marbacher Weg 6, D-35032, Marburg, Germany.
Organizational Affiliation: