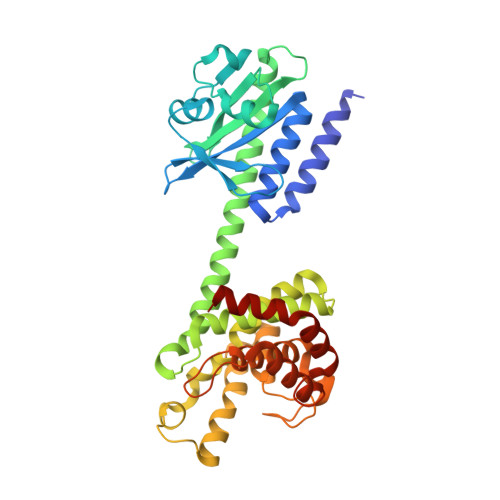
Crystal structure of an HD-GYP domain cyclic-di-GMP phosphodiesterase reveals an enzyme with a novel trinuclear catalytic iron centre.
Bellini, D., Caly, D.L., McCarthy, Y., Bumann, M., An, S.Q., Dow, J.M., Ryan, R.P., Walsh, M.A.(2014) Mol Microbiol 91: 26-38
- PubMed: 24176013 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/mmi.12447
- Primary Citation Related Structures:
4MCW, 4MDZ, 4ME4 - PubMed Abstract:
Bis-(3',5') cyclic di-guanylate (c-di-GMP) is a key bacterial second messenger that is implicated in the regulation of many crucial processes that include biofilm formation, motility and virulence. Cellular levels of c-di-GMP are controlled through synthesis by GGDEF domain diguanylate cyclases and degradation by two classes of phosphodiesterase with EAL or HD-GYP domains. Here, we have determined the structure of an enzymatically active HD-GYP domain protein from Persephonella marina (PmGH) alone, in complex with substrate (c-di-GMP) and final reaction product (GMP). The structures reveal a novel trinuclear iron binding site, which is implicated in catalysis and identify residues involved in recognition of c-di-GMP. This structure completes the picture of all domains involved in c-di-GMP metabolism and reveals that the HD-GYP family splits into two distinct subgroups containing bi- and trinuclear metal centres.
- Diamond Light Source, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0DE, UK; Research Complex at Harwell, Harwell Science and Innovation Campus, Didcot, Oxfordshire, OX11 0FA, UK.
Organizational Affiliation: