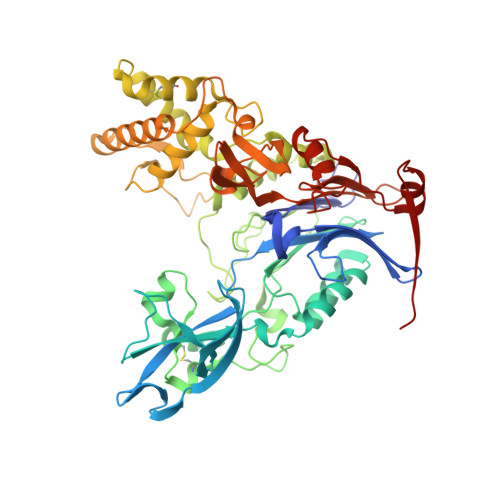
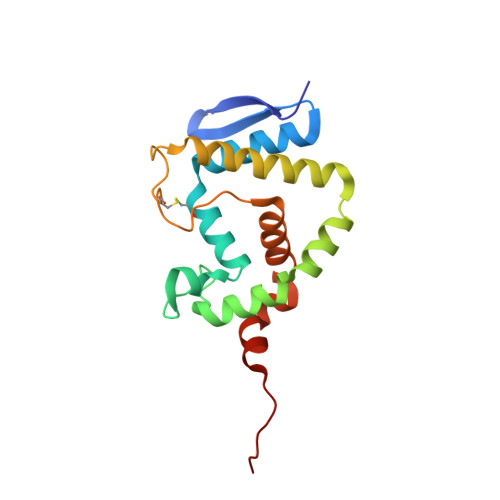
Rational Design of a Transition State Analogue with Picomolar Affinity for Pseudomonas aeruginosa PvdQ, a Siderophore Biosynthetic Enzyme.
Clevenger, K.D., Wu, R., Er, J.A., Liu, D., Fast, W.(2013) ACS Chem Biol 8: 2192-2200
- PubMed: 23883096 Search on PubMed
- DOI: https://doi.org/10.1021/cb400345h
- Primary Citation Related Structures:
4M1J - PubMed Abstract:
The Pseudomonas aeruginosa enzyme PvdQ can process different substrates involved in quorum-sensing or in siderophore biosynthesis. Substrate selectivity was evaluated using steady-state kinetic constants for hydrolysis of N-acyl-homoserine lactones (HSLs) and p-nitrophenyl fatty acid esters. PvdQ prefers substrates with alkyl chains between 12 and 14 carbons long that do not bear a 3-oxo substitution and is revealed here to have a relatively high specificity constant for selected N-acyl-HSLs (kcat/KM = 10(5) to 10(6) M(-1) s(-1)). However, endogenous P. aeruginosa N-acyl-HSLs are ≥100-fold disfavored, supporting the conclusion that PvdQ was not primarily evolved to regulate endogenous quorum-sensing. PvdQ plays an essential biosynthetic role for the siderophore pyoverdine, on which P. aeruginosa depends for growth in iron-limited environments. A series of alkylboronate inhibitors was found to be reversible, competitive, and extremely potent (Ki ≥ 190 pM). A 1.8 Å X-ray structure shows that 1-tridecylboronic acid forms a monocovalent bond with the N-terminal β-chain Ser residue in the PvdQ heterodimer, mimicking a reaction transition state. This boronic acid inhibits growth of P. aeruginosa in iron-limited media, reproducing the phenotype of a genetic pvdQ disruption, although co-administration of an efflux pump inhibitor is required to maintain growth inhibition. These findings support the strategy of designing boron-based inhibitors of siderophore biosynthetic enzymes to control P. aeruginosa infections.
- Department of Chemistry and Biochemistry and the ‡College of Pharmacy, Medicinal Chemistry Division, University of Texas , Austin, Texas 78712, United States.
Organizational Affiliation: