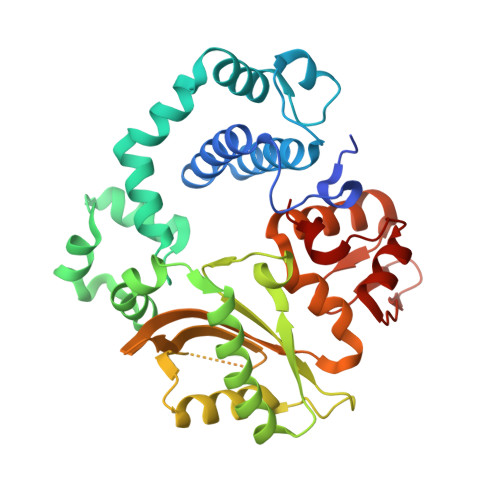
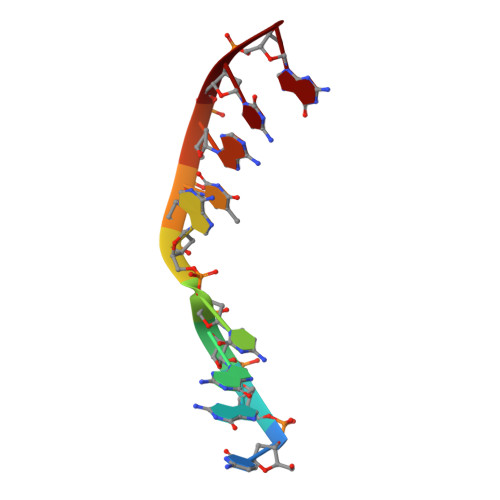
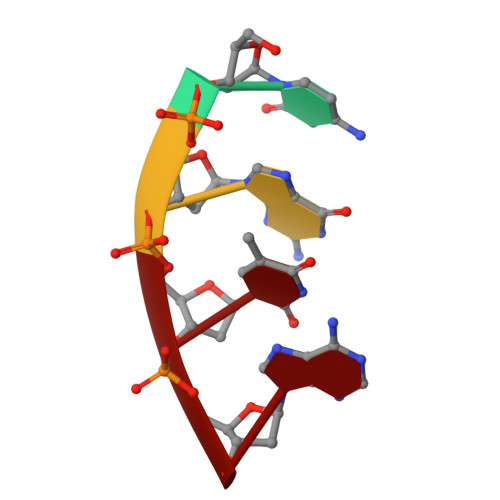
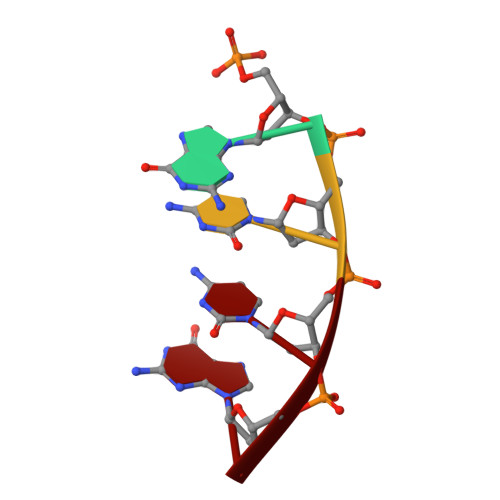
Sustained active site rigidity during synthesis by human DNA polymerase mu.
Moon, A.F., Pryor, J.M., Ramsden, D.A., Kunkel, T.A., Bebenek, K., Pedersen, L.C.(2014) Nat Struct Mol Biol 21: 253-260
- PubMed: 24487959 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.2766
- Primary Citation Related Structures:
4LZD, 4LZG, 4M04, 4M0A - PubMed Abstract:
DNA polymerase μ (Pol μ) is the only template-dependent human DNA polymerase capable of repairing double-strand DNA breaks (DSBs) with unpaired 3' ends in nonhomologous end joining (NHEJ). To probe this function, we structurally characterized Pol μ's catalytic cycle for single-nucleotide incorporation. These structures indicate that, unlike other template-dependent DNA polymerases, Pol μ shows no large-scale conformational changes in protein subdomains, amino acid side chains or DNA upon dNTP binding or catalysis. Instead, the only major conformational change is seen earlier in the catalytic cycle, when the flexible loop 1 region repositions upon DNA binding. Pol μ variants with changes in loop 1 have altered catalytic properties and are partially defective in NHEJ. The results indicate that specific loop 1 residues contribute to Pol μ's unique ability to catalyze template-dependent NHEJ of DSBs with unpaired 3' ends.
- Laboratory of Structural Biology, National Institute of Environmental Health Sciences, National Institutes of Health, Research Triangle Park, North Carolina, USA.
Organizational Affiliation: