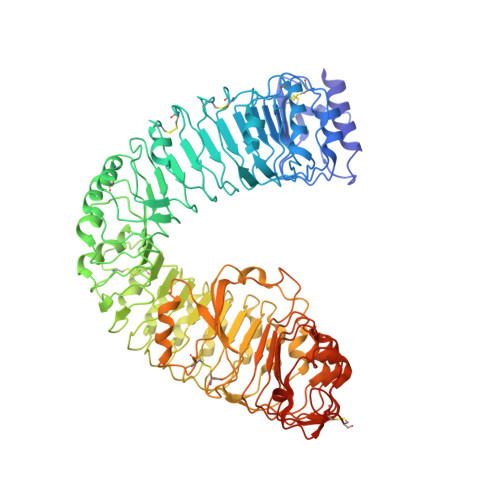
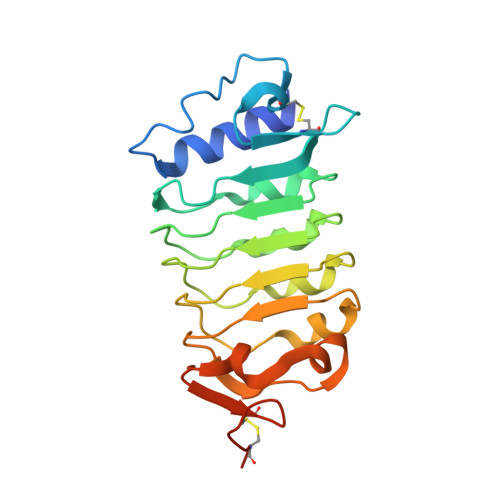
Molecular mechanism for plant steroid receptor activation by somatic embryogenesis co-receptor kinases.
Santiago, J., Henzler, C., Hothorn, M.(2013) Science 341: 889-892
- PubMed: 23929946 Search on PubMed
- DOI: https://doi.org/10.1126/science.1242468
- Primary Citation Related Structures:
4LSA, 4LSC, 4LSX - PubMed Abstract:
Brassinosteroids, which control plant growth and development, are sensed by the leucine-rich repeat (LRR) domain of the membrane receptor kinase BRASSINOSTEROID INSENSITIVE 1 (BRI1), but it is unknown how steroid binding at the cell surface activates the cytoplasmic kinase domain of the receptor. A family of somatic embryogenesis receptor kinases (SERKs) has been genetically implicated in mediating early brassinosteroid signaling events. We found a direct and steroid-dependent interaction between the BRI1 and SERK1 LRR domains by analysis of their complex crystal structure at 3.3 angstrom resolution. We show that the SERK1 LRR domain is involved in steroid sensing and, through receptor-co-receptor heteromerization, in the activation of the BRI1 signaling pathway. Our work reveals how known missense mutations in BRI1 and in SERKs modulate brassinosteroid signaling and the targeting mechanism of BRI1 receptor antagonists.
- Structural Plant Biology Lab, Friedrich Miescher Laboratory of the Max Planck Society, Spemannstraße 39, Tübingen 72076, Germany.
Organizational Affiliation: