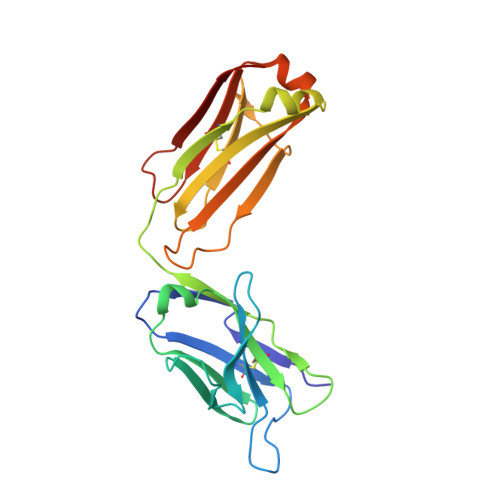
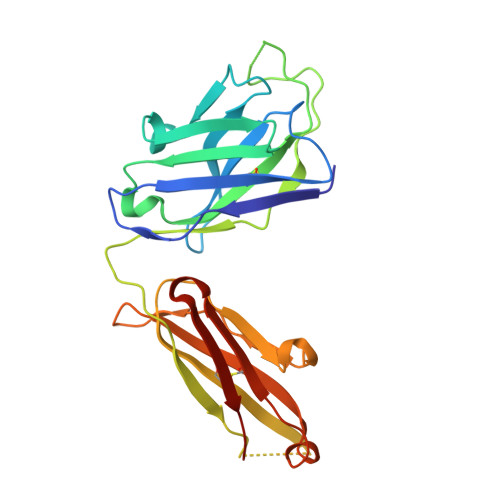
Mechanism for neutralizing activity by the anti-CMV gH/gL monoclonal antibody MSL-109.
Fouts, A.E., Comps-Agrar, L., Stengel, K.F., Ellerman, D., Schoeffler, A.J., Warming, S., Eaton, D.L., Feierbach, B.(2014) Proc Natl Acad Sci U S A 111: 8209-8214
- PubMed: 24843144 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1404653111
- Primary Citation Related Structures:
4LRI - PubMed Abstract:
Cytomegalovirus (CMV) is a widespread opportunistic pathogen that causes birth defects when transmitted transplacentally and severe systemic illness in immunocompromised individuals. MSL-109, a human monoclonal IgG isolated from a CMV seropositive individual, binds to the essential CMV entry glycoprotein H (gH) and prevents infection of cells. Here, we suggest a mechanism for neutralization activity by MSL-109. We define a genetic basis for resistance to MSL-109 and have generated a structural model of gH that reveals the epitope of this neutralizing antibody. Using surface-based, time-resolved FRET, we demonstrate that gH/gL interacts with glycoprotein B (gB). Additionally, we detect homodimers of soluble gH/gL heterodimers and confirm this novel oligomeric assembly on full-length gH/gL expressed on the cell surface. We show that MSL-109 perturbs the dimerization of gH/gL:gH/gL, suggesting that dimerization of gH/gL may be required for infectivity. gH/gL homodimerization may be conserved between alpha- and betaherpesviruses, because both CMV and HSV gH/gL demonstrate self-association in the FRET system. This study provides evidence for a novel mechanism of action for MSL-109 and reveals a previously undescribed aspect of viral entry that may be susceptible to therapeutic intervention.
- Departments of Infectious Diseases.
Organizational Affiliation: