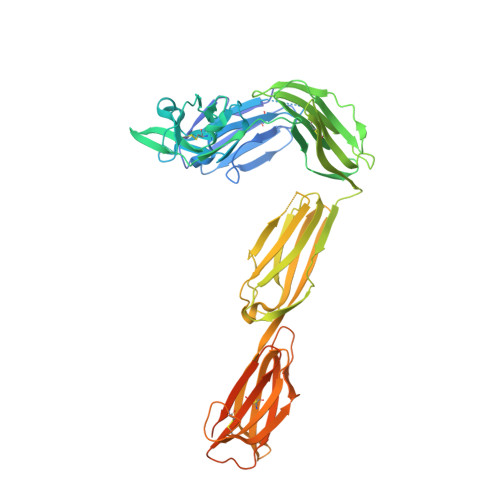
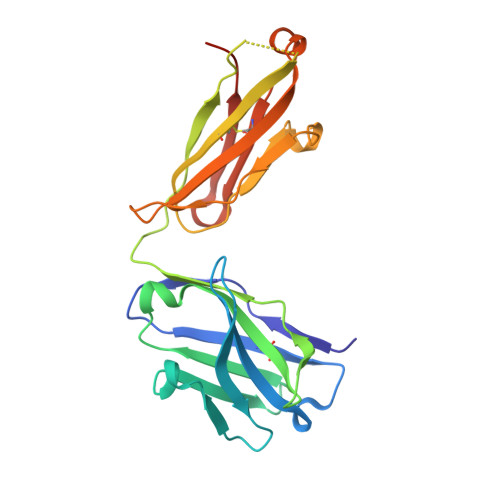
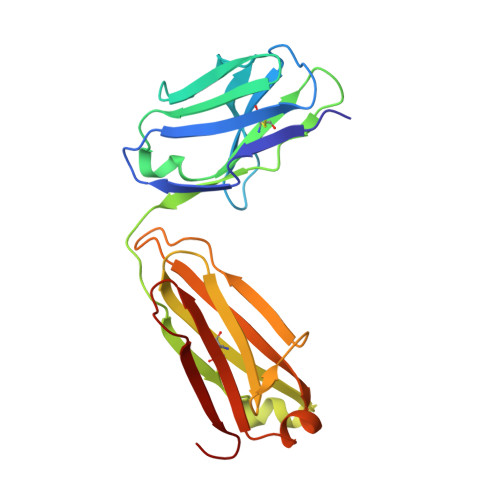
Targeting tumor-associated macrophages with anti-CSF-1R antibody reveals a strategy for cancer therapy
Ries, C.H., Cannarile, M.A., Hoves, S., Benz, J., Wartha, K., Runza, V., Rey-Giraud, F., Pradel, L.P., Feuerhake, F., Klaman, I., Jones, T., Jucknischke, U., Scheiblich, S., Kaluza, K., Gorr, I.H., Walz, A., Abiraj, K., Cassier, P.A., Sica, A., Gomez-Roca, C., de Visser, K.E., Italiano, A., Le Tourneau, C., Delord, J.P., Levitsky, H., Blay, J.Y., Ruttinger, D.(2014) Cancer Cell 25: 846-859
- PubMed: 24898549 Search on PubMed
- DOI: https://doi.org/10.1016/j.ccr.2014.05.016
- Primary Citation Related Structures:
4LIQ - PubMed Abstract:
Macrophage infiltration has been identified as an independent poor prognostic factor in several cancer types. The major survival factor for these macrophages is macrophage colony-stimulating factor 1 (CSF-1). We generated a monoclonal antibody (RG7155) that inhibits CSF-1 receptor (CSF-1R) activation. In vitro RG7155 treatment results in cell death of CSF-1-differentiated macrophages. In animal models, CSF-1R inhibition strongly reduces F4/80(+) tumor-associated macrophages accompanied by an increase of the CD8(+)/CD4(+) T cell ratio. Administration of RG7155 to patients led to striking reductions of CSF-1R(+)CD163(+) macrophages in tumor tissues, which translated into clinical objective responses in diffuse-type giant cell tumor (Dt-GCT) patients.
- Roche Innovation Center Penzberg, Oncology Division, Roche Pharmaceutical Research and Early Development, 82377 Penzberg, Germany. Electronic address: carola.ries@roche.com.
Organizational Affiliation: