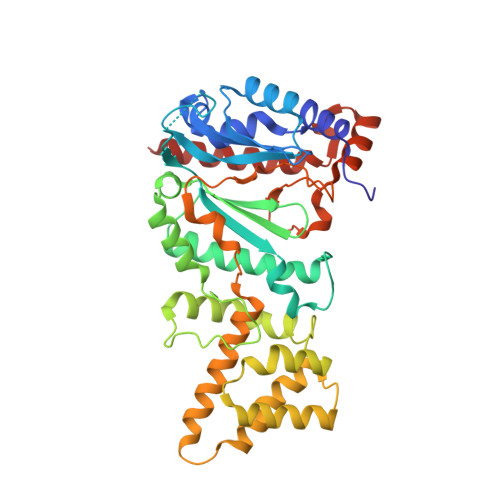
Insights into Eukaryotic Primer Synthesis from Structures of the p48 Subunit of Human DNA Primase.
Vaithiyalingam, S., Arnett, D.R., Aggarwal, A., Eichman, B.F., Fanning, E., Chazin, W.J.(2014) J Mol Biology 426: 558-569
- PubMed: 24239947 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2013.11.007
- Primary Citation Related Structures:
4LIK, 4LIL, 4LIM - PubMed Abstract:
DNA replication in all organisms requires polymerases to synthesize copies of the genome. DNA polymerases are unable to function on a bare template and require a primer. Primases are crucial RNA polymerases that perform the initial de novo synthesis, generating the first 8-10 nucleotides of the primer. Although structures of archaeal and bacterial primases have provided insights into general priming mechanisms, these proteins are not well conserved with heterodimeric (p48/p58) primases in eukaryotes. Here, we present X-ray crystal structures of the catalytic engine of a eukaryotic primase, which is contained in the p48 subunit. The structures of p48 reveal that eukaryotic primases maintain the conserved catalytic prim fold domain, but with a unique subdomain not found in the archaeal and bacterial primases. Calorimetry experiments reveal that Mn(2+) but not Mg(2+) significantly enhances the binding of nucleotide to primase, which correlates with higher catalytic efficiency in vitro. The structure of p48 with bound UTP and Mn(2+) provides insights into the mechanism of nucleotide synthesis by primase. Substitution of conserved residues involved in either metal or nucleotide binding alter nucleotide binding affinities, and yeast strains containing the corresponding Pri1p substitutions are not viable. Our results reveal that two residues (S160 and H166) in direct contact with the nucleotide were previously unrecognized as critical to the human primase active site. Comparing p48 structures to those of similar polymerases in different states of action suggests changes that would be required to attain a catalytically competent conformation capable of initiating dinucleotide synthesis.
- Department of Biochemistry, Vanderbilt University, Nashville, TN 37232, USA; Center for Structural Biology, Vanderbilt University, Nashville, TN 37232, USA.
Organizational Affiliation: