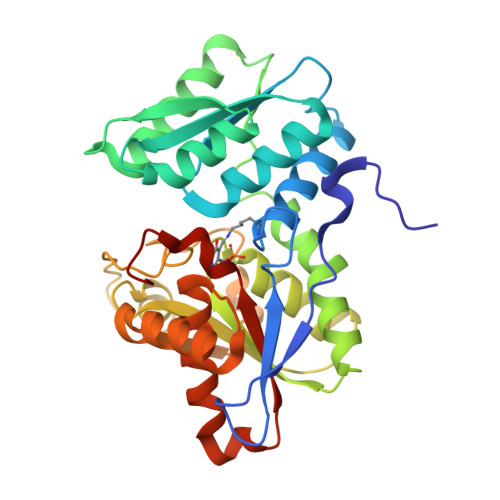
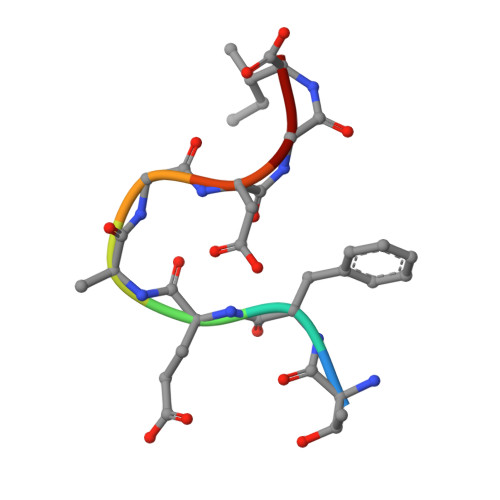
Crystal Structure of O-Acetylserine Sulfhydrylase from Haemophilus influenzae in complex with high affinity inhibitory peptide from Serine acetyl transferase of Salmonella typhimurium
Singh, A.K., Ekka, M.K., Kaushik, A., Kumaran, S.To be published.