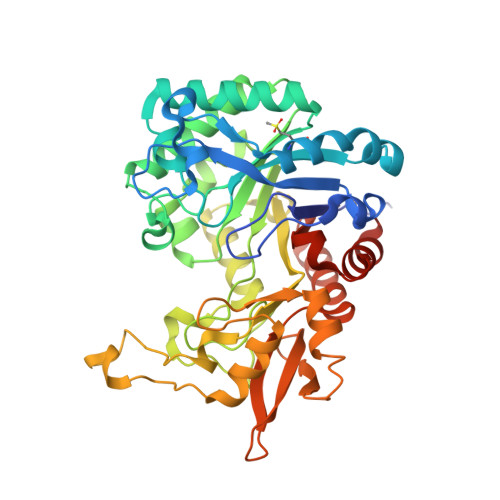
Inverse relationship between chitobiase and transglycosylation activities of chitinase-D from Serratia proteamaculans revealed by mutational and biophysical analyses.
Madhuprakash, J., Bobbili, K.B., Moerschbacher, B.M., Singh, T.P., Swamy, M.J., Podile, A.R.(2015) Sci Rep 5: 15657-15657
- PubMed: 26493546 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/srep15657
- Primary Citation Related Structures:
4LGX - PubMed Abstract:
Serratia proteamaculans chitinase-D (SpChiD) has a unique combination of hydrolytic and transglycosylation (TG) activities. The TG activity of SpChiD can be used for large-scale production of chito-oligosaccharides (CHOS). The multiple activities (hydrolytic and/or chitobiase activities and TG) of SpChiD appear to be strongly influenced by the substrate-binding cleft. Here, we report the unique property of SpChiD substrate-binding cleft, wherein, the residues Tyr28, Val35 and Thr36 control chitobiase activity and the residues Trp160 and Trp290 are crucial for TG activity. Mutants with reduced (V35G and T36G/F) or no (SpChiDΔ30-42 and Y28A) chitobiase activity produced higher amounts of the quantifiable even-chain TG product with degree of polymerization (DP)-6, indicating that the chitobiase and TG activities are inversely related. In addition to its unprecedented catalytic properties, unlike other chitinases, the single modular SpChiD showed dual unfolding transitions. Ligand-induced thermal stability studies with the catalytically inactive mutant of SpChiD (E153A) showed that the transition temperature increased upon binding of CHOS with DP2-6. Isothermal titration calorimetry experiments revealed the exceptionally high binding affinities for E153A to CHOS with DP2-6. These observations strongly support that the architecture of SpChiD substrate-binding cleft adopted to control chitobiase and TG activities, in addition to usual chitinase-mediated hydrolysis.
- Department of Plant Sciences, School of Life Sciences, University of Hyderabad, Gachibowli, Hyderabad, India.
Organizational Affiliation: