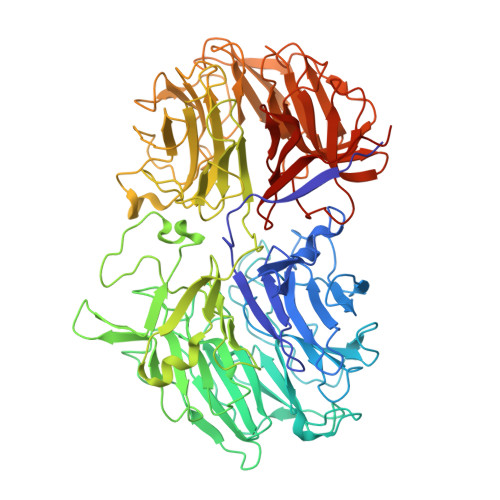
Structure of Acidothermus cellulolyticus family 74 glycoside hydrolase at 1.82 angstrom resolution.
Alahuhta, M., Adney, W.S., Himmel, M.E., Lunin, V.V.(2013) Acta Crystallogr Sect F Struct Biol Cryst Commun 69: 1335-1338
- PubMed: 24316824 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309113030005
- Primary Citation Related Structures:
4LGN - PubMed Abstract:
Here, a 1.82 Å resolution X-ray structure of a glycoside hydrolase family 74 (GH74) enzyme from Acidothermus cellulolyticus is reported. The resulting structure was refined to an R factor of 0.150 and an Rfree of 0.196. Structural analysis shows that five related structures have been reported with a secondary-structure similarity of between 75 and 89%. The five similar structures were all either Clostridium thermocellum or Geotrichum sp. M128 GH74 xyloglucanases. Structural analysis indicates that the A. cellulolyticus GH74 enzyme is an endoxyloglucanase, as it lacks a characteristic loop that blocks one end of the active site in exoxyloglucanases. Superimposition with the C. thermocellum GH74 shows that Asp451 and Asp38 are the catalytic residues.
- BioSciences Center, National Renewable Energy Laboratory, 15013 Denver West Parkway, Golden, CO 80401, USA.
Organizational Affiliation: