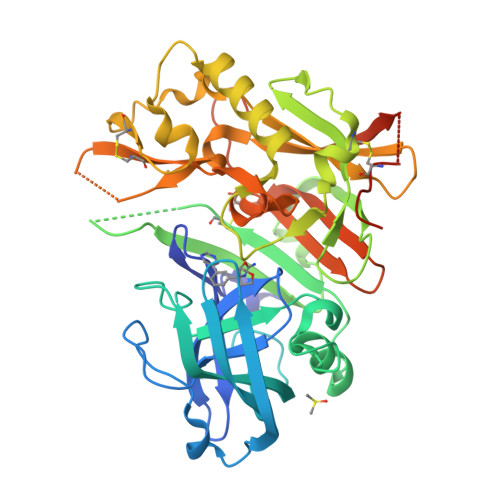
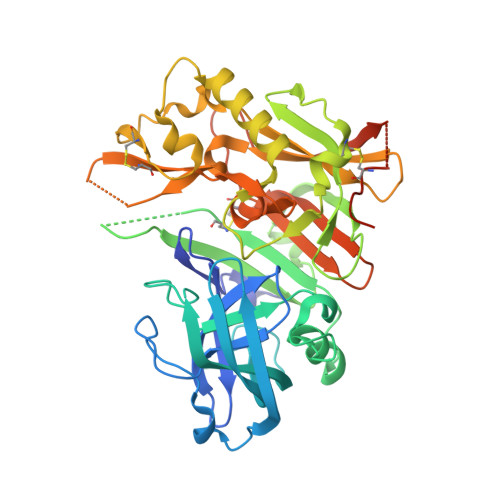
Diethylaminosulfur Trifluoride-Mediated Intramolecular Cyclization of 2-hydroxycycloalkylureas to Fused Bicyclic Aminooxazoline Compounds and Evaluation of Their Biochemical Activity Against β-Secretase-1 (BACE-1)
Huestis, M.P., Liu, W., Volgraf, M., Purkey, H., Yu, C., Wang, W., Smith, D., Vigers, G., Dutcher, D., Hunt, K.W., Siu, M.(2013) Tetrahedron Lett