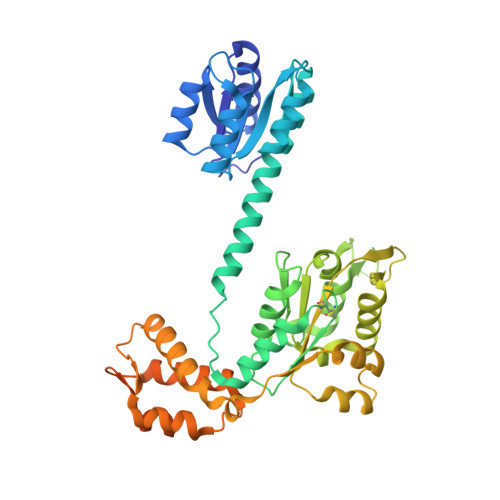
Structure, function, and tethering of DNA-binding domains in sigma (54) transcriptional activators.
Vidangos, N., Maris, A.E., Young, A., Hong, E., Pelton, J.G., Batchelor, J.D., Wemmer, D.E.(2013) Biopolymers 99: 1082-1096
- PubMed: 23818155 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/bip.22333
- Primary Citation Related Structures:
2M8G, 4L4U, 4L5E - PubMed Abstract:
We compare the structure, activity, and linkage of DNA-binding domains (DBDs) from σ(54) transcriptional activators and discuss how the properties of the DBDs and the linker to the neighboring domain are affected by the overall properties and requirements of the full proteins. These transcriptional activators bind upstream of specific promoters that utilize σ(54)-polymerase. Upon receiving a signal the activators assemble into hexamers, which then, through adenosine triphosphate (ATP) hydrolysis, drive a conformational change in polymerase that enables transcription initiation. We present structures of the DBDs of activators nitrogen regulatory protein C 1 (NtrC1) and Nif-like homolog 2 (Nlh2) from the thermophile Aquifex aeolicus. The structures of these domains and their relationship to other parts of the activators are discussed. These structures are compared with previously determined structures of the DBDs of NtrC4, NtrC, ZraR, and factor for inversion stimulation. The N-terminal linkers that connect the DBDs to the central domains in NtrC1 and Nlh2 were studied and found to be unstructured. Additionally, a crystal structure of full-length NtrC1 was solved, but density of the DBDs was extremely weak, further indicating that the linker between ATPase and DBDs functions as a flexible tether. Flexible linking of ATPase and DBDs is likely necessary to allow assembly of the active hexameric ATPase ring. The comparison of this set of activators also shows clearly that strong dimerization of the DBD only occurs when other domains do not dimerize strongly.
- Department of Chemistry and QB3 Institute, University of California, Berkeley, CA, 94720-1460.
Organizational Affiliation: