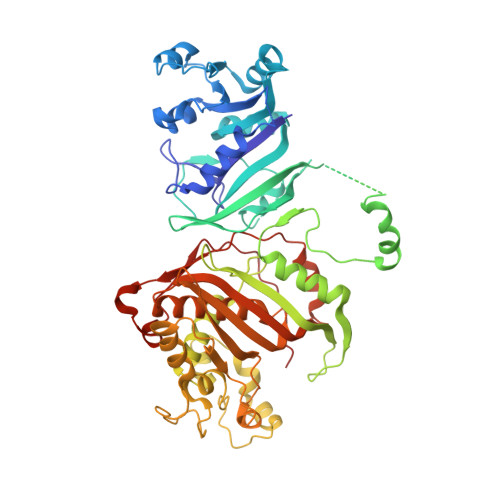
Discovery of potent and selective inhibitors of Toxoplasma gondii thymidylate synthase for opportunistic infections.
Zaware, N., Sharma, H., Yang, J., Devambatla, R.K., Queener, S.F., Anderson, K.S., Gangjee, A.(2013) ACS Med Chem Lett 4: 1148-1151
- PubMed: 24470841 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/ml400208v
- Primary Citation Related Structures:
4KY4, 4KYA - PubMed Abstract:
Infection by the parasite Toxoplasma gondii (tg) can lead to toxoplasmosis in immunocompromised patients such as organ transplant, cancer and HIV/AIDS patients. The bifunctional thymidylate synthase-dihydrofolate reductase (TS-DHFR) enzyme is crucial for nucleotide synthesis in T. gondii , and represents a potential target to combat T. gondii infection. While species selectivity with drugs has been attained for DHFR, TS is much more conserved across species and specificity is significantly more challenging. We discovered novel substituted-9 H -pyrimido[4,5- b ]indoles 1 - 3 with single-digit nanomolar K i for tgTS, two of which, 2 and 3 , are 28- and 122-fold selective over human TS (hTS). The synthesis of these compounds, and their structures in complex with tgTS-DHFR are presented along with binding measurements and cell culture data. These results show, for the very first time, that in spite of the high degree of conservation of active site residues between hTS and the parasite TS, specificity has been accomplished via novel structures and provides a new target (TS) for selective drug development against parasitic infections.
- Division of Medicinal Chemistry, Graduate School of Pharmaceutical Sciences, Duquesne University, 600 Forbes Avenue, Pittsburgh, PA 15282, United States.
Organizational Affiliation: