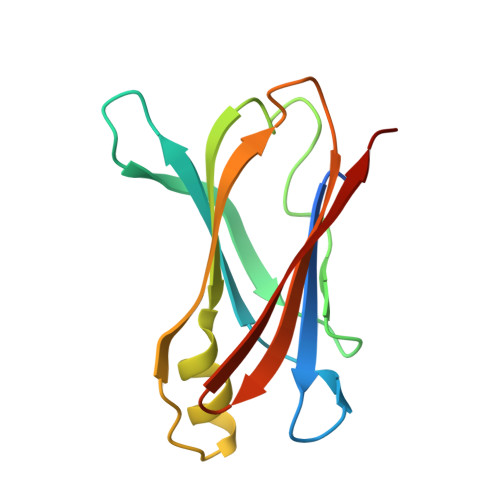
Bifunctional coumarin derivatives that inhibit transthyretin amyloidogenesis and serve as fluorescent transthyretin folding sensors.
Myung, N., Connelly, S., Kim, B., Park, S.J., Wilson, I.A., Kelly, J.W., Choi, S.(2013) Chem Commun (Camb) 49: 9188-9190
- PubMed: 23989101 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/c3cc44667k
- Primary Citation Related Structures:
4KY2 - PubMed Abstract:
We describe coumarin derivatives that are non-fluorescent in aqueous buffers and that very selectively bind to transthyretin (TTR) in complex biological environments potently inhibiting TTR amyloidogenesis while also exhibiting sensitive off-on fluorescent sensing of the properly folded quaternary structure of TTR.
- Department of New Drug Discovery and Development, Chungnam National University, Daejon, 305-764, Republic of Korea.
Organizational Affiliation: