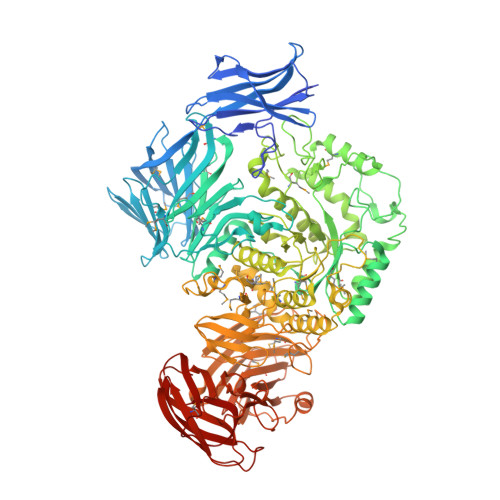
1.9 Angstrom resolution crystal structure of uncharacterized protein lmo2446 from Listeria monocytogenes EGD-e in complex with alpha-D-glucose, beta-D-glucose, magnesium and calcium
Halavaty, A.S., Minasov, G., Dubrovska, I., Winsor, J., Shuvalova, L., Peterson, S., Anderson, W.F., Center for Structural Genomics of Infectious Diseases (CSGID)To be published.