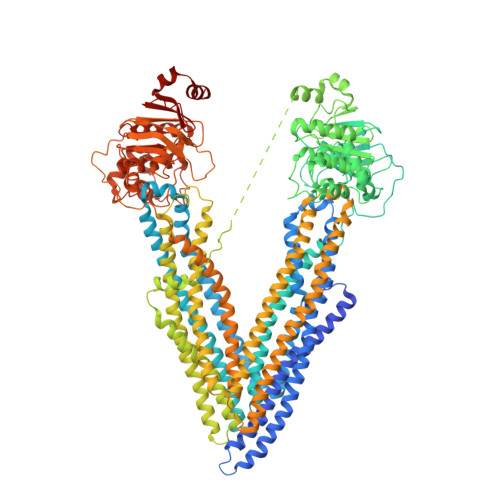
Structures of P-glycoprotein reveal its conformational flexibility and an epitope on the nucleotide-binding domain.
Ward, A.B., Szewczyk, P., Grimard, V., Lee, C.W., Martinez, L., Doshi, R., Caya, A., Villaluz, M., Pardon, E., Cregger, C., Swartz, D.J., Falson, P.G., Urbatsch, I.L., Govaerts, C., Steyaert, J., Chang, G.(2013) Proc Natl Acad Sci U S A 110: 13386-13391
- PubMed: 23901103 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1309275110
- Primary Citation Related Structures:
4KSB, 4KSC, 4KSD - PubMed Abstract:
P-glycoprotein (P-gp) is one of the best-known mediators of drug efflux-based multidrug resistance in many cancers. This validated therapeutic target is a prototypic, plasma membrane resident ATP-Binding Cassette transporter that pumps xenobiotic compounds out of cells. The large, polyspecific drug-binding pocket of P-gp recognizes a variety of structurally unrelated compounds. The transport of these drugs across the membrane is coincident with changes in the size and shape of this pocket during the course of the transport cycle. Here, we present the crystal structures of three inward-facing conformations of mouse P-gp derived from two different crystal forms. One structure has a nanobody bound to the C-terminal side of the first nucleotide-binding domain. This nanobody strongly inhibits the ATP hydrolysis activity of mouse P-gp by hindering the formation of a dimeric complex between the ATP-binding domains, which is essential for nucleotide hydrolysis. Together, these inward-facing conformational snapshots of P-gp demonstrate a range of flexibility exhibited by this transporter, which is likely an essential feature for the binding and transport of large, diverse substrates. The nanobody-bound structure also reveals a unique epitope on P-gp.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: