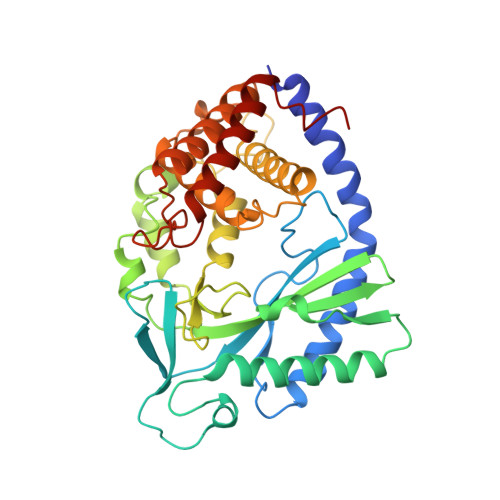
Structure of Human cGAS Reveals a Conserved Family of Second-Messenger Enzymes in Innate Immunity.
Kranzusch, P.J., Lee, A.S., Berger, J.M., Doudna, J.A.(2013) Cell Rep 3: 1362-1368
- PubMed: 23707061 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2013.05.008
- Primary Citation Related Structures:
4KM5 - PubMed Abstract:
Innate immune recognition of foreign nucleic acids induces protective interferon responses. Detection of cytosolic DNA triggers downstream immune signaling through activation of cyclic GMP-AMP synthase (cGAS). We report here the crystal structure of human cGAS, revealing an unanticipated zinc-ribbon DNA-binding domain appended to a core enzymatic nucleotidyltransferase scaffold. The catalytic core of cGAS is structurally homologous to the RNA-sensing enzyme, 2'-5' oligo-adenylate synthase (OAS), and divergent C-terminal domains account for specific ligand-activation requirements of each enzyme. We show that the cGAS zinc ribbon is essential for STING-dependent induction of the interferon response and that conserved amino acids displayed within the intervening loops are required for efficient cytosolic DNA recognition. These results demonstrate that cGAS and OAS define a family of innate immunity sensors and that structural divergence from a core nucleotidyltransferase enables second-messenger responses to distinct foreign nucleic acids.
- Department of Molecular & Cell Biology, Center for RNA Systems Biology, Howard Hughes Medical Institute, University of California, Berkeley, CA 94720, USA.
Organizational Affiliation: