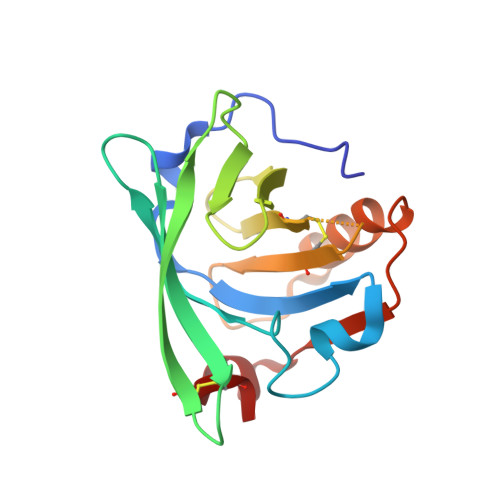
Structural Basis for Enantioselectivity in the Transfer Hydrogenation of a Ketone Catalyzed by an Artificial Metalloenzyme
Cherrier, M.V., Amara, P., Engilberge, S., Chevalley, A., Salmain, M., Fontecilla-Camps, J.C.(2013) Eur J Inorg Chem 21: 3596-3600