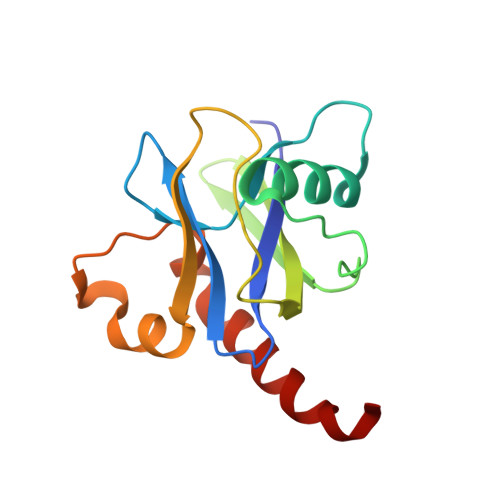
Active site conformational dynamics are coupled to catalysis in the mRNA decapping enzyme dcp2.
Aglietti, R.A., Floor, S.N., McClendon, C.L., Jacobson, M.P., Gross, J.D.(2013) Structure 21: 1571-1580
- PubMed: 23911090 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2013.06.021
- Primary Citation Related Structures:
4K6E, 4KG3, 4KG4 - PubMed Abstract:
Removal of the 5' cap structure by Dcp2 is a major step in several 5'-3' mRNA decay pathways. The activity of Dcp2 is enhanced by Dcp1 and bound coactivators, yet the details of how these interactions are linked to chemistry are poorly understood. Here, we report three crystal structures of the catalytic Nudix hydrolase domain of Dcp2 that demonstrate binding of a catalytically essential metal ion, and enzyme kinetics are used to identify several key active site residues involved in acid/base chemistry of decapping. Using nuclear magnetic resonance and molecular dynamics, we find that a conserved metal binding loop on the catalytic domain undergoes conformational changes during the catalytic cycle. These findings describe key events during the chemical step of decapping, suggest local active site conformational changes are important for activity, and provide a framework to explain stimulation of catalysis by the regulatory domain of Dcp2 and associated coactivators.
- Program in Chemistry and Chemical Biology, University of California, San Francisco, San Francisco, CA 94158, USA; Department of Pharmaceutical Chemistry, University of California, San Francisco, San Francisco, CA 94158, USA.
Organizational Affiliation: