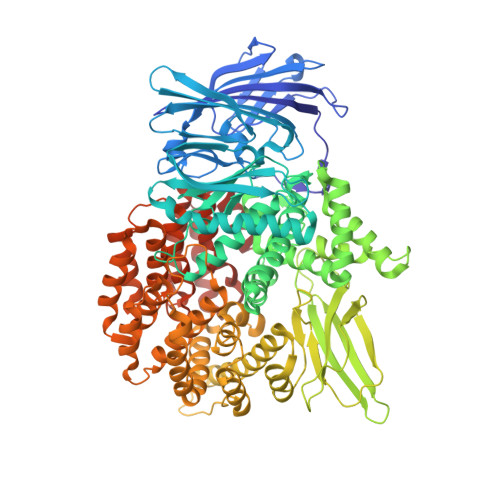
Synthesis and Structure-Activity Relationships of Phosphonic Arginine Mimetics as Inhibitors of the M1 and M17 Aminopeptidases from Plasmodium falciparum.
Kannan Sivaraman, K., Paiardini, A., Sienczyk, M., Ruggeri, C., Oellig, C.A., Dalton, J.P., Scammells, P.J., Drag, M., McGowan, S.(2013) J Med Chem 56: 5213-5217
- PubMed: 23713488 Search on PubMed
- DOI: https://doi.org/10.1021/jm4005972
- Primary Citation Related Structures:
4K3N, 4K5L, 4K5M, 4K5N, 4K5O, 4K5P - PubMed Abstract:
The malaria parasite Plasmodium falciparum employs two metallo-aminopeptidases, PfA-M1 and PfA-M17, which are essential for parasite survival. Compounds that inhibit the activity of either enzyme represent leads for the development of new antimalarial drugs. Here we report the synthesis and structure-activity relationships of a small library of phosphonic acid arginine mimetics that probe the S1 pocket of both enzymes and map the necessary interactions that would be important for a dual inhibitor.
- Department of Biochemistry and Molecular Biology, Monash University, Clayton Campus, Melbourne, VIC 3800, Australia.
Organizational Affiliation: