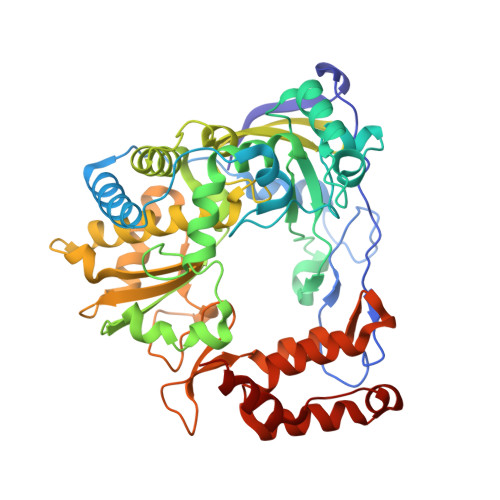
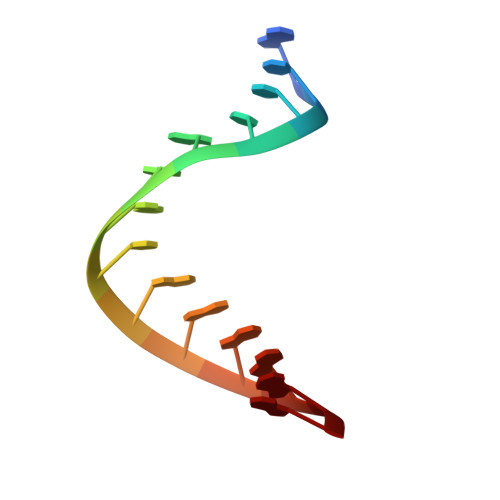
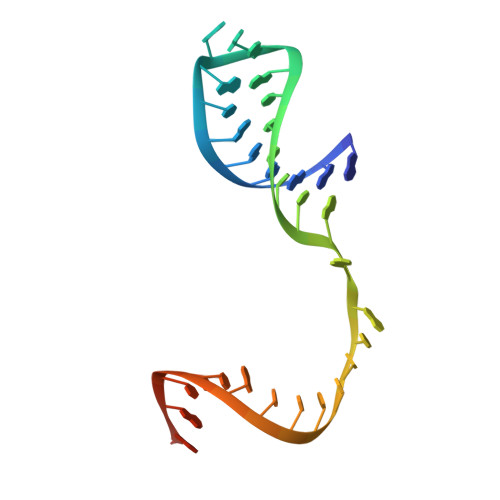Structures of coxsackievirus, rhinovirus, and poliovirus polymerase elongation complexes solved by engineering RNA mediated crystal contacts.
Gong, P., Kortus, M.G., Nix, J.C., Davis, R.E., Peersen, O.B.(2013) PLoS One 8: e60272-e60272
- PubMed: 23667424 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0060272
- Primary Citation Related Structures:
4K4S, 4K4T, 4K4U, 4K4V, 4K4W, 4K4X, 4K4Y, 4K4Z, 4K50 - PubMed Abstract:
RNA-dependent RNA polymerases play a vital role in the growth of RNA viruses where they are responsible for genome replication, but do so with rather low fidelity that allows for the rapid adaptation to different host cell environments. These polymerases are also a target for antiviral drug development. However, both drug discovery efforts and our understanding of fidelity determinants have been hampered by a lack of detailed structural information about functional polymerase-RNA complexes and the structural changes that take place during the elongation cycle. Many of the molecular details associated with nucleotide selection and catalysis were revealed in our recent structure of the poliovirus polymerase-RNA complex solved by first purifying and then crystallizing stalled elongation complexes. In the work presented here we extend that basic methodology to determine nine new structures of poliovirus, coxsackievirus, and rhinovirus elongation complexes at 2.2-2.9 Å resolution. The structures highlight conserved features of picornaviral polymerases and the interactions they make with the template and product RNA strands, including a tight grip on eight basepairs of the nascent duplex, a fully pre-positioned templating nucleotide, and a conserved binding pocket for the +2 position template strand base. At the active site we see a pre-bound magnesium ion and there is conservation of a non-standard backbone conformation of the template strand in an interaction that may aid in triggering RNA translocation via contact with the conserved polymerase motif B. Moreover, by engineering plasticity into RNA-RNA contacts, we obtain crystal forms that are capable of multiple rounds of in-crystal catalysis and RNA translocation. Together, the data demonstrate that engineering flexible RNA contacts to promote crystal lattice formation is a versatile platform that can be used to solve the structures of viral RdRP elongation complexes and their catalytic cycle intermediates.
- Department of Biochemistry and Molecular Biology, Colorado State University, Fort Collins, Colorado, United States of America.
Organizational Affiliation: