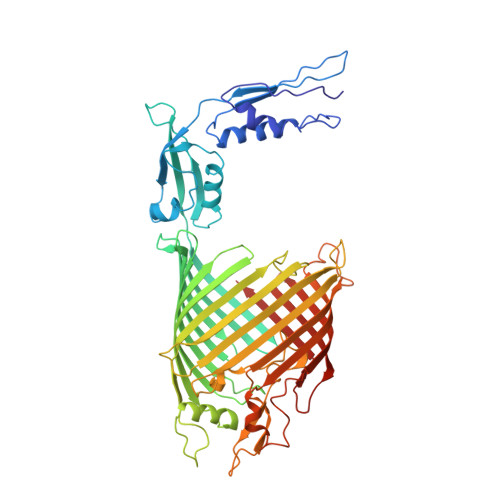
Structural insight into the biogenesis of beta-barrel membrane proteins.
Noinaj, N., Kuszak, A.J., Gumbart, J.C., Lukacik, P., Chang, H., Easley, N.C., Lithgow, T., Buchanan, S.K.(2013) Nature 501: 385-390
- PubMed: 23995689 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nature12521
- Primary Citation Related Structures:
4K3B, 4K3C - PubMed Abstract:
β-barrel membrane proteins are essential for nutrient import, signalling, motility and survival. In Gram-negative bacteria, the β-barrel assembly machinery (BAM) complex is responsible for the biogenesis of β-barrel membrane proteins, with homologous complexes found in mitochondria and chloroplasts. Here we describe the structure of BamA, the central and essential component of the BAM complex, from two species of bacteria: Neisseria gonorrhoeae and Haemophilus ducreyi. BamA consists of a large periplasmic domain attached to a 16-strand transmembrane β-barrel domain. Three structural features shed light on the mechanism by which BamA catalyses β-barrel assembly. First, the interior cavity is accessible in one BamA structure and conformationally closed in the other. Second, an exterior rim of the β-barrel has a distinctly narrowed hydrophobic surface, locally destabilizing the outer membrane. And third, the β-barrel can undergo lateral opening, suggesting a route from the interior cavity in BamA into the outer membrane.
- National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, Maryland 20892, USA.
Organizational Affiliation: