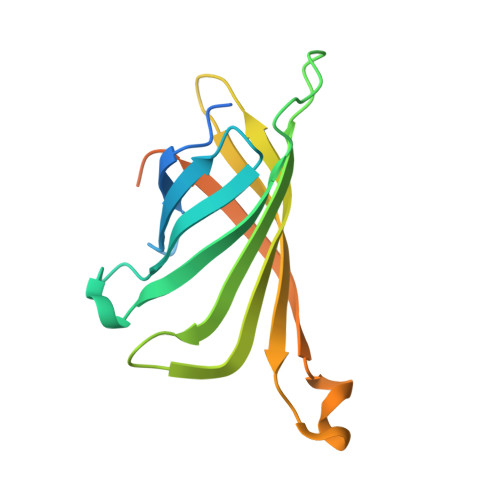
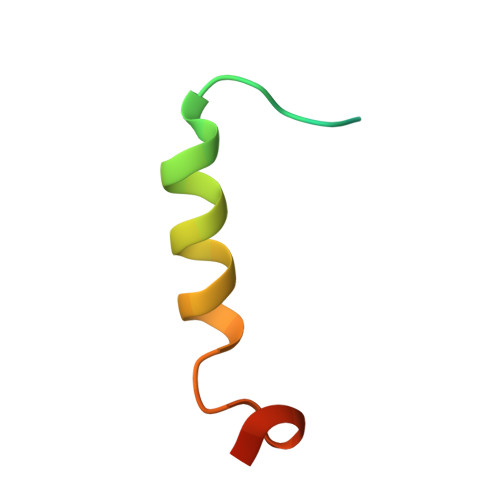
The structure of the SBP-Tag-streptavidin complex reveals a novel helical scaffold bridging binding pockets on separate subunits
Barrette-Ng, I.H., Wu, S.C., Tjia, W.M., Wong, S.L., Ng, K.K.(2013) Acta Crystallogr D Biol Crystallogr 69: 879-887
- PubMed: 23633599 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S0907444913002576
- Primary Citation Related Structures:
4JO6 - PubMed Abstract:
The 38-residue SBP-Tag binds to streptavidin more tightly (K(d) -/= 2.5-4.9 nM) than most if not all other known peptide sequences. Crystallographic analysis at 1.75 Å resolution shows that the SBP-Tag binds to streptavidin in an unprecedented manner by simultaneously interacting with biotin-binding pockets from two separate subunits. An N-terminal HVV peptide sequence (residues 12-14) and a C-terminal HPQ sequence (residues 31-33) form the bulk of the direct interactions between the SBP-Tag and the two biotin-binding pockets. Surprisingly, most of the peptide spanning these two sites (residues 17-28) adopts a regular α-helical structure that projects three leucine side chains into a groove formed at the interface between two streptavidin protomers. The crystal structure shows that residues 1-10 and 35-38 of the original SBP-Tag identified through in vitro selection and deletion analysis do not appear to contact streptavidin and thus may not be important for binding. A 25-residue peptide comprising residues 11-34 (SBP-Tag2) was synthesized and shown using surface plasmon resonance to bind streptavidin with very similar affinity and kinetics when compared with the SBP-Tag. The SBP-Tag2 was also added to the C-terminus of β-lactamase and was shown to be just as effective as the full-length SBP-Tag in affinity purification. These results validate the molecular structure of the SBP-Tag-streptavidin complex and establish a minimal bivalent streptavidin-binding tag from which further rational design and optimization can proceed.
- Department of Biological Sciences, University of Calgary, 2500 University Drive NW, Calgary, Alberta T2N 1N4, Canada.
Organizational Affiliation: